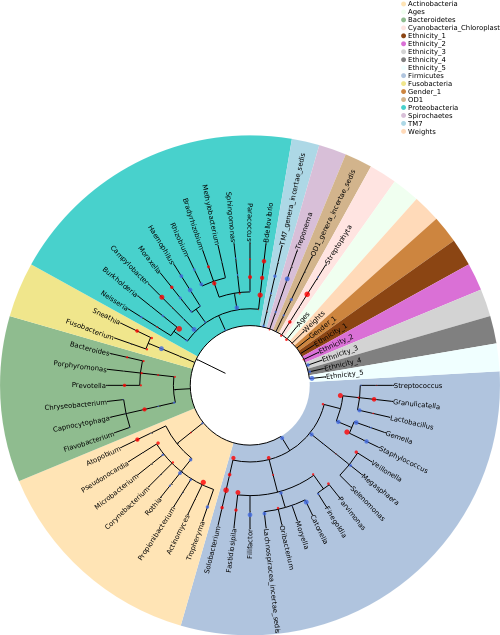
Example : k fold cross-validation with an input file¶
DeepBiome package takes microbiome abundance data as input and uses the phylogenetic taxonomy to guide the decision of the optimal number of layers and neurons in the deep learning architecture.
To use DeepBiome, you can experiment (1) k times repetition or (2) k fold cross-validation. For each experiment, we asuume that the dataset is given by - A list of k input files for k times repetition. - One input file for k fold cross-validation.
This notebook contains an example of (2) k fold cross-validation for the deep neural netowrk using deepbiome.
1. Load library¶
First, we load the DeepBiome package. The DeepBiome package is built on the tensorflow and keras library
[1]:
import os
import logging
import json
from pkg_resources import resource_filename
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
pd.set_option('precision', 3)
pd.set_option('chop_threshold', .0005)
np.set_printoptions(precision=3, suppress=True)
from deepbiome import deepbiome
Using TensorFlow backend.
2. Prepare the dataset¶
In this example, we assume that we have one input file for k times repetition.
DeepBiome needs 3 data files as follows: 1. the tree information 1. the input file 1. y
In addition, we can set the training index for each fold. If we set the index file, DeepBiome builds the training set for each fold based on each fold index in the index file. If not, DeepBiome will generate the index file locally.
Each data should have the csv format as follow:
Example of the tree information¶
First we need a file about the phylogenetic tree information. This tree information file should have the format below:
[2]:
tree_information = pd.read_csv(resource_filename('deepbiome', 'tests/data/genus48_dic.csv'))
tree_information
[2]:
| Genus | Family | Order | Class | Phylum | |
|---|---|---|---|---|---|
| 0 | Streptococcus | Streptococcaceae | Lactobacillales | Bacilli | Firmicutes |
| 1 | Tropheryma | Cellulomonadaceae | Actinomycetales | Actinobacteria | Actinobacteria |
| 2 | Veillonella | Veillonellaceae | Selenomonadales | Negativicutes | Firmicutes |
| 3 | Actinomyces | Actinomycetaceae | Actinomycetales | Actinobacteria | Actinobacteria |
| 4 | Flavobacterium | Flavobacteriaceae | Flavobacteriales | Flavobacteria | Bacteroidetes |
| 5 | Prevotella | Prevotellaceae | Bacteroidales | Bacteroidia | Bacteroidetes |
| 6 | Porphyromonas | Porphyromonadaceae | Bacteroidales | Bacteroidia | Bacteroidetes |
| 7 | Parvimonas | Clostridiales_Incertae_Sedis_XI | Clostridiales | Clostridia | Firmicutes |
| 8 | Fusobacterium | Fusobacteriaceae | Fusobacteriales | Fusobacteria | Fusobacteria |
| 9 | Propionibacterium | Propionibacteriaceae | Actinomycetales | Actinobacteria | Actinobacteria |
| 10 | Gemella | Bacillales_Incertae_Sedis_XI | Bacillales | Bacilli | Firmicutes |
| 11 | Rothia | Micrococcaceae | Actinomycetales | Actinobacteria | Actinobacteria |
| 12 | Granulicatella | Carnobacteriaceae | Lactobacillales | Bacilli | Firmicutes |
| 13 | Neisseria | Neisseriaceae | Neisseriales | Betaproteobacteria | Proteobacteria |
| 14 | Lactobacillus | Lactobacillaceae | Lactobacillales | Bacilli | Firmicutes |
| 15 | Megasphaera | Veillonellaceae | Selenomonadales | Negativicutes | Firmicutes |
| 16 | Catonella | Lachnospiraceae | Clostridiales | Clostridia | Firmicutes |
| 17 | Atopobium | Coriobacteriaceae | Coriobacteriales | Actinobacteria | Actinobacteria |
| 18 | Campylobacter | Campylobacteraceae | Campylobacterales | Epsilonproteobacteria | Proteobacteria |
| 19 | Capnocytophaga | Flavobacteriaceae | Flavobacteriales | Flavobacteria | Bacteroidetes |
| 20 | Solobacterium | Erysipelotrichaceae | Erysipelotrichales | Erysipelotrichia | Firmicutes |
| 21 | Moryella | Lachnospiraceae | Clostridiales | Clostridia | Firmicutes |
| 22 | TM7_genera_incertae_sedis | TM7_genera_incertae_sedis | TM7_genera_incertae_sedis | TM7_genera_incertae_sedis | TM7 |
| 23 | Staphylococcus | Staphylococcaceae | Bacillales | Bacilli | Firmicutes |
| 24 | Filifactor | Peptostreptococcaceae | Clostridiales | Clostridia | Firmicutes |
| 25 | Oribacterium | Lachnospiraceae | Clostridiales | Clostridia | Firmicutes |
| 26 | Burkholderia | Burkholderiaceae | Burkholderiales | Betaproteobacteria | Proteobacteria |
| 27 | Sneathia | Leptotrichiaceae | Fusobacteriales | Fusobacteria | Fusobacteria |
| 28 | Treponema | Spirochaetaceae | Spirochaetales | Spirochaetes | Spirochaetes |
| 29 | Moraxella | Moraxellaceae | Pseudomonadales | Gammaproteobacteria | Proteobacteria |
| 30 | Haemophilus | Pasteurellaceae | Pasteurellales | Gammaproteobacteria | Proteobacteria |
| 31 | Selenomonas | Veillonellaceae | Selenomonadales | Negativicutes | Firmicutes |
| 32 | Corynebacterium | Corynebacteriaceae | Actinomycetales | Actinobacteria | Actinobacteria |
| 33 | Rhizobium | Rhizobiaceae | Rhizobiales | Alphaproteobacteria | Proteobacteria |
| 34 | Bradyrhizobium | Bradyrhizobiaceae | Rhizobiales | Alphaproteobacteria | Proteobacteria |
| 35 | Methylobacterium | Methylobacteriaceae | Rhizobiales | Alphaproteobacteria | Proteobacteria |
| 36 | OD1_genera_incertae_sedis | OD1_genera_incertae_sedis | OD1_genera_incertae_sedis | OD1_genera_incertae_sedis | OD1 |
| 37 | Finegoldia | Clostridiales_Incertae_Sedis_XI | Clostridiales | Clostridia | Firmicutes |
| 38 | Microbacterium | Microbacteriaceae | Actinomycetales | Actinobacteria | Actinobacteria |
| 39 | Sphingomonas | Sphingomonadaceae | Sphingomonadales | Alphaproteobacteria | Proteobacteria |
| 40 | Chryseobacterium | Flavobacteriaceae | Flavobacteriales | Flavobacteria | Bacteroidetes |
| 41 | Bacteroides | Bacteroidaceae | Bacteroidales | Bacteroidia | Bacteroidetes |
| 42 | Bdellovibrio | Bdellovibrionaceae | Bdellovibrionales | Deltaproteobacteria | Proteobacteria |
| 43 | Streptophyta | Chloroplast | Chloroplast | Chloroplast | Cyanobacteria_Chloroplast |
| 44 | Lachnospiracea_incertae_sedis | Lachnospiraceae | Clostridiales | Clostridia | Firmicutes |
| 45 | Paracoccus | Rhodobacteraceae | Rhodobacterales | Alphaproteobacteria | Proteobacteria |
| 46 | Fastidiosipila | Ruminococcaceae | Clostridiales | Clostridia | Firmicutes |
| 47 | Pseudonocardia | Pseudonocardiaceae | Actinomycetales | Actinobacteria | Actinobacteria |
This file has .csv format below:
[3]:
with open(resource_filename('deepbiome', 'tests/data/genus48_dic.csv')) as f:
print(f.read())
Genus,Family,Order,Class,Phylum
Streptococcus,Streptococcaceae,Lactobacillales,Bacilli,Firmicutes
Tropheryma,Cellulomonadaceae,Actinomycetales,Actinobacteria,Actinobacteria
Veillonella,Veillonellaceae,Selenomonadales,Negativicutes,Firmicutes
Actinomyces,Actinomycetaceae,Actinomycetales,Actinobacteria,Actinobacteria
Flavobacterium,Flavobacteriaceae,Flavobacteriales,Flavobacteria,Bacteroidetes
Prevotella,Prevotellaceae,Bacteroidales,Bacteroidia,Bacteroidetes
Porphyromonas,Porphyromonadaceae,Bacteroidales,Bacteroidia,Bacteroidetes
Parvimonas,Clostridiales_Incertae_Sedis_XI,Clostridiales,Clostridia,Firmicutes
Fusobacterium,Fusobacteriaceae,Fusobacteriales,Fusobacteria,Fusobacteria
Propionibacterium,Propionibacteriaceae,Actinomycetales,Actinobacteria,Actinobacteria
Gemella,Bacillales_Incertae_Sedis_XI,Bacillales,Bacilli,Firmicutes
Rothia,Micrococcaceae,Actinomycetales,Actinobacteria,Actinobacteria
Granulicatella,Carnobacteriaceae,Lactobacillales,Bacilli,Firmicutes
Neisseria,Neisseriaceae,Neisseriales,Betaproteobacteria,Proteobacteria
Lactobacillus,Lactobacillaceae,Lactobacillales,Bacilli,Firmicutes
Megasphaera,Veillonellaceae,Selenomonadales,Negativicutes,Firmicutes
Catonella,Lachnospiraceae,Clostridiales,Clostridia,Firmicutes
Atopobium,Coriobacteriaceae,Coriobacteriales,Actinobacteria,Actinobacteria
Campylobacter,Campylobacteraceae,Campylobacterales,Epsilonproteobacteria,Proteobacteria
Capnocytophaga,Flavobacteriaceae,Flavobacteriales,Flavobacteria,Bacteroidetes
Solobacterium,Erysipelotrichaceae,Erysipelotrichales,Erysipelotrichia,Firmicutes
Moryella,Lachnospiraceae,Clostridiales,Clostridia,Firmicutes
TM7_genera_incertae_sedis,TM7_genera_incertae_sedis,TM7_genera_incertae_sedis,TM7_genera_incertae_sedis,TM7
Staphylococcus,Staphylococcaceae,Bacillales,Bacilli,Firmicutes
Filifactor,Peptostreptococcaceae,Clostridiales,Clostridia,Firmicutes
Oribacterium,Lachnospiraceae,Clostridiales,Clostridia,Firmicutes
Burkholderia,Burkholderiaceae,Burkholderiales,Betaproteobacteria,Proteobacteria
Sneathia,Leptotrichiaceae,Fusobacteriales,Fusobacteria,Fusobacteria
Treponema,Spirochaetaceae,Spirochaetales,Spirochaetes,Spirochaetes
Moraxella,Moraxellaceae,Pseudomonadales,Gammaproteobacteria,Proteobacteria
Haemophilus,Pasteurellaceae,Pasteurellales,Gammaproteobacteria,Proteobacteria
Selenomonas,Veillonellaceae,Selenomonadales,Negativicutes,Firmicutes
Corynebacterium,Corynebacteriaceae,Actinomycetales,Actinobacteria,Actinobacteria
Rhizobium,Rhizobiaceae,Rhizobiales,Alphaproteobacteria,Proteobacteria
Bradyrhizobium,Bradyrhizobiaceae,Rhizobiales,Alphaproteobacteria,Proteobacteria
Methylobacterium,Methylobacteriaceae,Rhizobiales,Alphaproteobacteria,Proteobacteria
OD1_genera_incertae_sedis,OD1_genera_incertae_sedis,OD1_genera_incertae_sedis,OD1_genera_incertae_sedis,OD1
Finegoldia,Clostridiales_Incertae_Sedis_XI,Clostridiales,Clostridia,Firmicutes
Microbacterium,Microbacteriaceae,Actinomycetales,Actinobacteria,Actinobacteria
Sphingomonas,Sphingomonadaceae,Sphingomonadales,Alphaproteobacteria,Proteobacteria
Chryseobacterium,Flavobacteriaceae,Flavobacteriales,Flavobacteria,Bacteroidetes
Bacteroides,Bacteroidaceae,Bacteroidales,Bacteroidia,Bacteroidetes
Bdellovibrio,Bdellovibrionaceae,Bdellovibrionales,Deltaproteobacteria,Proteobacteria
Streptophyta,Chloroplast,Chloroplast,Chloroplast,Cyanobacteria_Chloroplast
Lachnospiracea_incertae_sedis,Lachnospiraceae,Clostridiales,Clostridia,Firmicutes
Paracoccus,Rhodobacteraceae,Rhodobacterales,Alphaproteobacteria,Proteobacteria
Fastidiosipila,Ruminococcaceae,Clostridiales,Clostridia,Firmicutes
Pseudonocardia,Pseudonocardiaceae,Actinomycetales,Actinobacteria,Actinobacteria
Example of the input file¶
Below is an example of the input file. This example has 1000 samples’ microbiome abandunce. The order of the microbiome should be same as the order of the microbiome in the Genus level in the tree information above.
[4]:
x = pd.read_csv(resource_filename('deepbiome', 'tests/data/onefile_x.csv'))
x.head()
[4]:
| Streptococcus | Tropheryma | Veillonella | Actinomyces | Flavobacterium | Prevotella | Porphyromonas | Parvimonas | Fusobacterium | Propionibacterium | ... | Microbacterium | Sphingomonas | Chryseobacterium | Bacteroides | Bdellovibrio | Streptophyta | Lachnospiracea_incertae_sedis | Paracoccus | Fastidiosipila | Pseudonocardia | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 0 | 841 | 0 | 813 | 505 | 5 | 3224 | 0 | 362 | 11 | 65 | ... | 0 | 87 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 2133 |
| 1 | 1445 | 0 | 1 | 573 | 0 | 1278 | 82 | 85 | 69 | 154 | ... | 0 | 1 | 2 | 0 | 0 | 0 | 0 | 0 | 0 | 3638 |
| 2 | 1259 | 0 | 805 | 650 | 0 | 1088 | 0 | 0 | 74 | 0 | ... | 0 | 2 | 8 | 1 | 39 | 0 | 0 | 0 | 0 | 3445 |
| 3 | 982 | 0 | 327 | 594 | 0 | 960 | 81 | 19 | 9 | 0 | ... | 157 | 1 | 0 | 4 | 60 | 0 | 0 | 0 | 0 | 3507 |
| 4 | 1162 | 0 | 130 | 969 | 163 | 1515 | 167 | 4 | 162 | 3 | ... | 0 | 9 | 0 | 0 | 0 | 0 | 60 | 0 | 0 | 3945 |
5 rows × 48 columns
[5]:
x.tail()
[5]:
| Streptococcus | Tropheryma | Veillonella | Actinomyces | Flavobacterium | Prevotella | Porphyromonas | Parvimonas | Fusobacterium | Propionibacterium | ... | Microbacterium | Sphingomonas | Chryseobacterium | Bacteroides | Bdellovibrio | Streptophyta | Lachnospiracea_incertae_sedis | Paracoccus | Fastidiosipila | Pseudonocardia | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 995 | 1401 | 4 | 30 | 526 | 0 | 923 | 25 | 0 | 127 | 0 | ... | 0 | 0 | 7 | 0 | 0 | 0 | 0 | 0 | 0 | 4470 |
| 996 | 2655 | 6 | 106 | 74 | 0 | 952 | 76 | 13 | 158 | 125 | ... | 0 | 2 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 2826 |
| 997 | 335 | 0 | 71 | 259 | 67 | 718 | 1 | 4 | 4 | 167 | ... | 0 | 246 | 0 | 0 | 6 | 0 | 0 | 0 | 0 | 6527 |
| 998 | 649 | 69 | 966 | 1227 | 0 | 508 | 2 | 30 | 550 | 0 | ... | 0 | 0 | 0 | 0 | 0 | 6 | 0 | 0 | 0 | 4402 |
| 999 | 1258 | 0 | 0 | 1119 | 0 | 2348 | 25 | 0 | 137 | 176 | ... | 0 | 2 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 2585 |
5 rows × 48 columns
This file has .csv format below:
[6]:
with open(resource_filename('deepbiome', 'tests/data/onefile_x.csv')) as f:
x_csv = f.readlines()
_ = [print(l) for l in x_csv[:10]]
"Streptococcus","Tropheryma","Veillonella","Actinomyces","Flavobacterium","Prevotella","Porphyromonas","Parvimonas","Fusobacterium","Propionibacterium","Gemella","Rothia","Granulicatella","Neisseria","Lactobacillus","Megasphaera","Catonella","Atopobium","Campylobacter","Capnocytophaga","Solobacterium","Moryella","TM7_genera_incertae_sedis","Staphylococcus","Filifactor","Oribacterium","Burkholderia","Sneathia","Treponema","Moraxella","Haemophilus","Selenomonas","Corynebacterium","Rhizobium","Bradyrhizobium","Methylobacterium","OD1_genera_incertae_sedis","Finegoldia","Microbacterium","Sphingomonas","Chryseobacterium","Bacteroides","Bdellovibrio","Streptophyta","Lachnospiracea_incertae_sedis","Paracoccus","Fastidiosipila","Pseudonocardia"
841,0,813,505,5,3224,0,362,11,65,156,1,55,0,1,20,382,1,333,24,80,43,309,2,3,4,0,1,32,0,2,4,382,0,0,96,23,0,0,87,0,0,0,0,0,0,0,2133
1445,0,1,573,0,1278,82,85,69,154,436,3,0,61,440,0,394,83,33,123,0,49,414,0,0,37,0,0,42,0,0,384,27,0,0,0,146,0,0,1,2,0,0,0,0,0,0,3638
1259,0,805,650,0,1088,0,0,74,0,155,228,430,765,0,0,11,102,68,90,77,83,322,10,0,7,0,122,76,0,1,25,0,0,0,44,13,0,0,2,8,1,39,0,0,0,0,3445
982,0,327,594,0,960,81,19,9,0,45,457,1049,0,3,450,19,170,388,147,0,0,41,63,0,1,0,0,121,0,0,1,0,0,0,0,344,0,157,1,0,4,60,0,0,0,0,3507
1162,0,130,969,163,1515,167,4,162,3,12,0,48,73,93,259,52,0,201,85,14,14,434,2,0,0,0,0,187,0,0,188,45,0,0,0,4,0,0,9,0,0,0,0,60,0,0,3945
1956,37,41,661,47,1555,374,7,142,19,61,226,0,30,52,0,6,480,142,148,9,575,12,0,0,0,0,3,168,0,56,50,0,0,0,98,989,0,0,12,0,0,0,0,0,0,0,2044
1037,14,83,1595,132,305,103,174,1195,0,410,224,1,320,26,0,476,0,7,37,46,61,20,0,0,0,0,0,226,0,239,8,1,0,0,0,0,188,0,20,4,0,4,0,0,0,0,3044
641,0,172,179,0,1312,84,9,81,376,128,223,160,0,532,155,89,355,1,282,0,0,25,0,0,43,0,9,311,0,0,0,0,0,0,0,845,0,0,8,0,0,0,0,0,0,0,3980
852,146,504,99,2,376,116,152,67,0,120,3,23,2,34,0,127,75,240,60,42,0,9,0,15,0,62,0,13,0,197,187,396,0,0,20,51,0,0,3,0,0,0,0,0,0,0,6007
Example of the Y (regression)¶
This is an example of the output file for regression problem.
[7]:
y = pd.read_csv(resource_filename('deepbiome', 'tests/data/onefile_regression_y.csv'))
y.head()
[7]:
| x1 | |
|---|---|
| 0 | 4.997 |
| 1 | 5.004 |
| 2 | 5.485 |
| 3 | 5.490 |
| 4 | 1.500 |
[8]:
y.tail()
[8]:
| x1 | |
|---|---|
| 995 | 2.610 |
| 996 | 5.489 |
| 997 | 3.498 |
| 998 | 5.486 |
| 999 | 5.320 |
Example of the Y (classification)¶
This is an example of the output file for classification problem. Below example file has 1000 samples in rows.
[9]:
y = pd.read_csv(resource_filename('deepbiome', 'tests/data/onefile_classification_y.csv'))
y.head()
[9]:
| V1 | |
|---|---|
| 0 | 1.000 |
| 1 | 1.000 |
| 2 | 0.000 |
| 3 | 0.000 |
| 4 | 1.000 |
[10]:
y.tail()
[10]:
| V1 | |
|---|---|
| 995 | 1.000 |
| 996 | 0.000 |
| 997 | 1.000 |
| 998 | 0.000 |
| 999 | 1.000 |
Exmple of the training index file for k fold cross-validation¶
For each fold, we have to set the training and test set. If the index file is given, DeepBiome sets the training set and test set based on the index file for 5 fold cross-validation. Below is the example of the index file. Each column has the training indices for each fold. DeepBiome will only use the samples in this index set for training.
[11]:
idxs = pd.read_csv(resource_filename('deepbiome', 'tests/data/onefile_idx.csv'), dtype=np.int)
idxs.head()
[11]:
| 0 | 1 | 2 | 3 | 4 | |
|---|---|---|---|---|---|
| 0 | 1 | 1 | 3 | 1 | 1 |
| 1 | 2 | 2 | 4 | 2 | 2 |
| 2 | 3 | 3 | 5 | 3 | 4 |
| 3 | 6 | 4 | 7 | 4 | 5 |
| 4 | 7 | 5 | 9 | 5 | 6 |
[12]:
idxs.tail()
[12]:
| 0 | 1 | 2 | 3 | 4 | |
|---|---|---|---|---|---|
| 795 | 994 | 996 | 994 | 995 | 996 |
| 796 | 995 | 997 | 995 | 996 | 997 |
| 797 | 997 | 998 | 996 | 997 | 998 |
| 798 | 999 | 999 | 998 | 998 | 999 |
| 799 | 1000 | 1000 | 999 | 1000 | 1000 |
Below is the index set for 1st fold. From 1000 samples above, it uses 800 samples for training.
[13]:
idxs.iloc[:,0].head()
[13]:
0 1
1 2
2 3
3 6
4 7
Name: 0, dtype: int64
[14]:
idxs.iloc[:,0].tail()
[14]:
795 994
796 995
797 997
798 999
799 1000
Name: 0, dtype: int64
3. Prepare the configuration¶
For detailed configuration, we can build the configuration information for the network training by: 1. the python dictionary format 1. the configufation file (.cfg).
In this notebook, we show the python dictionary format configuration.
Please check the detailed information about each option in the documantation
For preparing the configuration about the network information (network_info)¶
To give the information about the training process, we provide a dictionary of configurations to the netowrk_info field. Your configuration for the network training should include the information about:
[15]:
network_info = {
'architecture_info': {
'batch_normalization': 'False',
'drop_out': '0',
'weight_initial': 'glorot_uniform',
'weight_l1_penalty':'0.',
'weight_decay': 'phylogenetic_tree',
},
'model_info': {
'lr': '0.01',
'decay': '0.001',
'loss': 'binary_crossentropy',
'metrics': 'binary_accuracy, sensitivity, specificity, gmeasure',
'taxa_selection_metrics': 'sensitivity, specificity, gmeasure, accuracy',
'network_class': 'DeepBiomeNetwork',
'optimizer': 'adam',
'reader_class': 'MicroBiomeClassificationReader',
'normalizer': 'normalize_minmax',
},
'training_info': {
'epochs': '10',
'batch_size': '50',
'callbacks': 'ModelCheckpoint',
'monitor': 'val_loss',
'mode' : 'min',
'min_delta': '1e-7',
},
'validation_info': {
'batch_size': 'None',
'validation_size': '0.2'
},
'test_info': {
'batch_size': 'None'
}
}
For preparing the configuration about the path information (path_info)¶
To give the information about the path of dataset, paths for saving the trained weights and the evaluation results, we provide a dictionary of configurations to the path_info field. Your configuration for the network training should include the information about:
[16]:
path_info = {
'data_info': {
'data_path': resource_filename('deepbiome', 'tests/data'),
'idx_path': resource_filename('deepbiome', 'tests/data/onefile_idx.csv'),
'tree_info_path': resource_filename('deepbiome', 'tests/data/genus48_dic.csv'),
'x_path': 'onefile_x.csv',
'y_path': 'classification_y.csv'
},
'model_info': {
'evaluation': 'eval.npy',
'history': 'hist.json',
'model_dir': './example_result/',
'weight': 'weight.h5'
}
}
4. Deepbiome Training¶
Now we can train the DeepBiome network based on the configurations.
For logging, we used the python logging library.
[17]:
logging.basicConfig(format = '[%(name)-8s|%(levelname)s|%(filename)s:%(lineno)s] %(message)s',
level=logging.DEBUG)
log = logging.getLogger()
The deeobiome_train function provide the test evaluation, train evaluation and the deepbiome network instance.
If we set number_of_fold, then DeepBiome performs cross-validation based on that value. If not, DeepBiome package performs cross-validation based on the index file. If both number_of_fold option and the index file are missing, then the library performs leave-one-out-cross-validation (LOOCV).
[18]:
test_evaluation, train_evaluation, network = deepbiome.deepbiome_train(log, network_info, path_info, number_of_fold=None)
[root |INFO|deepbiome.py:115] -----------------------------------------------------------------
[root |INFO|deepbiome.py:153] -------1 simulation start!----------------------------------
[root |INFO|readers.py:58] -----------------------------------------------------------------------
[root |INFO|readers.py:59] Construct Dataset
[root |INFO|readers.py:60] -----------------------------------------------------------------------
[root |INFO|readers.py:61] Load data
[root |INFO|deepbiome.py:164] -----------------------------------------------------------------
[root |INFO|deepbiome.py:165] Build network for 1 simulation
[root |INFO|build_network.py:521] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:522] Read phylogenetic tree information from /DATA/home/muha/github_repos/deepbiome/deepbiome/tests/data/genus48_dic.csv
[root |INFO|build_network.py:528] Phylogenetic tree level list: ['Genus', 'Family', 'Order', 'Class', 'Phylum']
[root |INFO|build_network.py:529] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:537] Genus: 48
[root |INFO|build_network.py:537] Family: 40
[root |INFO|build_network.py:537] Order: 23
[root |INFO|build_network.py:537] Class: 17
[root |INFO|build_network.py:537] Phylum: 9
[root |INFO|build_network.py:546] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:547] Phylogenetic_tree_dict info: ['Genus', 'Order', 'Class', 'Family', 'Phylum', 'Number']
[root |INFO|build_network.py:548] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:558] Build edge weights between [ Genus, Family]
[root |INFO|build_network.py:558] Build edge weights between [Family, Order]
[root |INFO|build_network.py:558] Build edge weights between [ Order, Class]
[root |INFO|build_network.py:558] Build edge weights between [ Class, Phylum]
[root |INFO|build_network.py:571] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:586] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:587] Build network based on phylogenetic tree information
[root |INFO|build_network.py:588] ------------------------------------------------------------------------------------------
WARNING:tensorflow:From /usr/local/lib/python3.5/dist-packages/tensorflow/python/ops/resource_variable_ops.py:432: colocate_with (from tensorflow.python.framework.ops) is deprecated and will be removed in a future version.
Instructions for updating:
Colocations handled automatically by placer.
[tensorflow|WARNING|deprecation.py:328] From /usr/local/lib/python3.5/dist-packages/tensorflow/python/ops/resource_variable_ops.py:432: colocate_with (from tensorflow.python.framework.ops) is deprecated and will be removed in a future version.
Instructions for updating:
Colocations handled automatically by placer.
[root |INFO|build_network.py:670] ------------------------------------------------------------------------------------------
Model: "model_1"
_________________________________________________________________
Layer (type) Output Shape Param #
=================================================================
input (InputLayer) (None, 48) 0
_________________________________________________________________
l1_dense (Dense_with_tree) (None, 40) 1960
_________________________________________________________________
l1_activation (Activation) (None, 40) 0
_________________________________________________________________
l2_dense (Dense_with_tree) (None, 23) 943
_________________________________________________________________
l2_activation (Activation) (None, 23) 0
_________________________________________________________________
l3_dense (Dense_with_tree) (None, 17) 408
_________________________________________________________________
l3_activation (Activation) (None, 17) 0
_________________________________________________________________
l4_dense (Dense_with_tree) (None, 9) 162
_________________________________________________________________
l4_activation (Activation) (None, 9) 0
_________________________________________________________________
last_dense_h (Dense) (None, 1) 10
_________________________________________________________________
p_hat (Activation) (None, 1) 0
=================================================================
Total params: 3,483
Trainable params: 3,483
Non-trainable params: 0
_________________________________________________________________
[root |INFO|build_network.py:61] Build Network
[root |INFO|build_network.py:62] Optimizer = adam
[root |INFO|build_network.py:63] Loss = binary_crossentropy
[root |INFO|build_network.py:64] Metrics = binary_accuracy, sensitivity, specificity, gmeasure
[root |INFO|deepbiome.py:176] -----------------------------------------------------------------
[root |INFO|deepbiome.py:177] 1 fold computing start!----------------------------------
[root |INFO|build_network.py:137] Training start!
WARNING:tensorflow:From /usr/local/lib/python3.5/dist-packages/tensorflow/python/ops/math_ops.py:2862: to_int32 (from tensorflow.python.ops.math_ops) is deprecated and will be removed in a future version.
Instructions for updating:
Use tf.cast instead.
[tensorflow|WARNING|deprecation.py:328] From /usr/local/lib/python3.5/dist-packages/tensorflow/python/ops/math_ops.py:2862: to_int32 (from tensorflow.python.ops.math_ops) is deprecated and will be removed in a future version.
Instructions for updating:
Use tf.cast instead.
Train on 640 samples, validate on 160 samples
Epoch 1/10
640/640 [==============================] - 1s 1ms/step - loss: 0.6650 - binary_accuracy: 0.6625 - sensitivity: 0.9644 - specificity: 0.0427 - gmeasure: 0.0420 - val_loss: 0.6235 - val_binary_accuracy: 0.6938 - val_sensitivity: 1.0000 - val_specificity: 0.0000e+00 - val_gmeasure: 0.0000e+00
Epoch 2/10
640/640 [==============================] - 0s 134us/step - loss: 0.6316 - binary_accuracy: 0.6844 - sensitivity: 1.0000 - specificity: 0.0000e+00 - gmeasure: 0.0000e+00 - val_loss: 0.6170 - val_binary_accuracy: 0.6938 - val_sensitivity: 1.0000 - val_specificity: 0.0000e+00 - val_gmeasure: 0.0000e+00
Epoch 3/10
640/640 [==============================] - 0s 134us/step - loss: 0.6259 - binary_accuracy: 0.6844 - sensitivity: 1.0000 - specificity: 0.0000e+00 - gmeasure: 0.0000e+00 - val_loss: 0.6176 - val_binary_accuracy: 0.6938 - val_sensitivity: 1.0000 - val_specificity: 0.0000e+00 - val_gmeasure: 0.0000e+00
Epoch 4/10
640/640 [==============================] - 0s 139us/step - loss: 0.6239 - binary_accuracy: 0.6844 - sensitivity: 1.0000 - specificity: 0.0000e+00 - gmeasure: 0.0000e+00 - val_loss: 0.6162 - val_binary_accuracy: 0.6938 - val_sensitivity: 1.0000 - val_specificity: 0.0000e+00 - val_gmeasure: 0.0000e+00
Epoch 5/10
640/640 [==============================] - 0s 152us/step - loss: 0.6237 - binary_accuracy: 0.6844 - sensitivity: 1.0000 - specificity: 0.0000e+00 - gmeasure: 0.0000e+00 - val_loss: 0.6161 - val_binary_accuracy: 0.6938 - val_sensitivity: 1.0000 - val_specificity: 0.0000e+00 - val_gmeasure: 0.0000e+00
Epoch 6/10
640/640 [==============================] - 0s 161us/step - loss: 0.6246 - binary_accuracy: 0.6844 - sensitivity: 1.0000 - specificity: 0.0000e+00 - gmeasure: 0.0000e+00 - val_loss: 0.6169 - val_binary_accuracy: 0.6938 - val_sensitivity: 1.0000 - val_specificity: 0.0000e+00 - val_gmeasure: 0.0000e+00
Epoch 7/10
640/640 [==============================] - 0s 153us/step - loss: 0.6237 - binary_accuracy: 0.6844 - sensitivity: 1.0000 - specificity: 0.0000e+00 - gmeasure: 0.0000e+00 - val_loss: 0.6161 - val_binary_accuracy: 0.6938 - val_sensitivity: 1.0000 - val_specificity: 0.0000e+00 - val_gmeasure: 0.0000e+00
Epoch 8/10
640/640 [==============================] - 0s 150us/step - loss: 0.6249 - binary_accuracy: 0.6844 - sensitivity: 1.0000 - specificity: 0.0000e+00 - gmeasure: 0.0000e+00 - val_loss: 0.6161 - val_binary_accuracy: 0.6938 - val_sensitivity: 1.0000 - val_specificity: 0.0000e+00 - val_gmeasure: 0.0000e+00
Epoch 9/10
640/640 [==============================] - 0s 145us/step - loss: 0.6247 - binary_accuracy: 0.6844 - sensitivity: 1.0000 - specificity: 0.0000e+00 - gmeasure: 0.0000e+00 - val_loss: 0.6174 - val_binary_accuracy: 0.6938 - val_sensitivity: 1.0000 - val_specificity: 0.0000e+00 - val_gmeasure: 0.0000e+00
Epoch 10/10
640/640 [==============================] - 0s 137us/step - loss: 0.6237 - binary_accuracy: 0.6844 - sensitivity: 1.0000 - specificity: 0.0000e+00 - gmeasure: 0.0000e+00 - val_loss: 0.6159 - val_binary_accuracy: 0.6938 - val_sensitivity: 1.0000 - val_specificity: 0.0000e+00 - val_gmeasure: 0.0000e+00
[root |INFO|build_network.py:87] Load trained model weight at ./example_result/weight_0.h5
[root |INFO|build_network.py:147] Training end with time 4.15680456161499!
[root |INFO|build_network.py:83] Saved trained model weight at ./example_result/weight_0.h5
[root |DEBUG|deepbiome.py:185] Save weight at ./example_result/weight_0.h5
[root |DEBUG|deepbiome.py:188] Save history at ./example_result/hist_0.json
[root |INFO|build_network.py:173] Evaluation start!
800/800 [==============================] - 0s 3us/step
[root |INFO|build_network.py:178] Evaluation end with time 0.009856224060058594!
[root |INFO|build_network.py:179] Evaluation: [0.6216976642608643, 0.6862499713897705, 1.0, 0.0, 0.0]
[root |INFO|build_network.py:173] Evaluation start!
200/200 [==============================] - 0s 10us/step
[root |INFO|build_network.py:178] Evaluation end with time 0.008182048797607422!
[root |INFO|build_network.py:179] Evaluation: [0.6188781261444092, 0.6899999976158142, 1.0, 0.0, 0.0]
[root |INFO|deepbiome.py:199] Compute time : 5.385685920715332
[root |INFO|deepbiome.py:200] 1 fold computing end!---------------------------------------------
[root |INFO|deepbiome.py:153] -------2 simulation start!----------------------------------
[root |INFO|readers.py:58] -----------------------------------------------------------------------
[root |INFO|readers.py:59] Construct Dataset
[root |INFO|readers.py:60] -----------------------------------------------------------------------
[root |INFO|readers.py:61] Load data
[root |INFO|deepbiome.py:164] -----------------------------------------------------------------
[root |INFO|deepbiome.py:165] Build network for 2 simulation
[root |INFO|build_network.py:521] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:522] Read phylogenetic tree information from /DATA/home/muha/github_repos/deepbiome/deepbiome/tests/data/genus48_dic.csv
[root |INFO|build_network.py:528] Phylogenetic tree level list: ['Genus', 'Family', 'Order', 'Class', 'Phylum']
[root |INFO|build_network.py:529] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:537] Genus: 48
[root |INFO|build_network.py:537] Family: 40
[root |INFO|build_network.py:537] Order: 23
[root |INFO|build_network.py:537] Class: 17
[root |INFO|build_network.py:537] Phylum: 9
[root |INFO|build_network.py:546] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:547] Phylogenetic_tree_dict info: ['Genus', 'Order', 'Class', 'Family', 'Phylum', 'Number']
[root |INFO|build_network.py:548] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:558] Build edge weights between [ Genus, Family]
[root |INFO|build_network.py:558] Build edge weights between [Family, Order]
[root |INFO|build_network.py:558] Build edge weights between [ Order, Class]
[root |INFO|build_network.py:558] Build edge weights between [ Class, Phylum]
[root |INFO|build_network.py:571] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:586] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:587] Build network based on phylogenetic tree information
[root |INFO|build_network.py:588] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:670] ------------------------------------------------------------------------------------------
Model: "model_1"
_________________________________________________________________
Layer (type) Output Shape Param #
=================================================================
input (InputLayer) (None, 48) 0
_________________________________________________________________
l1_dense (Dense_with_tree) (None, 40) 1960
_________________________________________________________________
l1_activation (Activation) (None, 40) 0
_________________________________________________________________
l2_dense (Dense_with_tree) (None, 23) 943
_________________________________________________________________
l2_activation (Activation) (None, 23) 0
_________________________________________________________________
l3_dense (Dense_with_tree) (None, 17) 408
_________________________________________________________________
l3_activation (Activation) (None, 17) 0
_________________________________________________________________
l4_dense (Dense_with_tree) (None, 9) 162
_________________________________________________________________
l4_activation (Activation) (None, 9) 0
_________________________________________________________________
last_dense_h (Dense) (None, 1) 10
_________________________________________________________________
p_hat (Activation) (None, 1) 0
=================================================================
Total params: 3,483
Trainable params: 3,483
Non-trainable params: 0
_________________________________________________________________
[root |INFO|build_network.py:61] Build Network
[root |INFO|build_network.py:62] Optimizer = adam
[root |INFO|build_network.py:63] Loss = binary_crossentropy
[root |INFO|build_network.py:64] Metrics = binary_accuracy, sensitivity, specificity, gmeasure
[root |INFO|deepbiome.py:176] -----------------------------------------------------------------
[root |INFO|deepbiome.py:177] 2 fold computing start!----------------------------------
[root |INFO|build_network.py:137] Training start!
Train on 640 samples, validate on 160 samples
Epoch 1/10
640/640 [==============================] - 0s 768us/step - loss: 0.6595 - binary_accuracy: 0.7078 - sensitivity: 0.9801 - specificity: 0.0121 - gmeasure: 0.0263 - val_loss: 0.6135 - val_binary_accuracy: 0.7375 - val_sensitivity: 1.0000 - val_specificity: 0.0000e+00 - val_gmeasure: 0.0000e+00
Epoch 2/10
640/640 [==============================] - 0s 127us/step - loss: 0.6079 - binary_accuracy: 0.7156 - sensitivity: 1.0000 - specificity: 0.0000e+00 - gmeasure: 0.0000e+00 - val_loss: 0.5757 - val_binary_accuracy: 0.7375 - val_sensitivity: 1.0000 - val_specificity: 0.0000e+00 - val_gmeasure: 0.0000e+00
Epoch 3/10
640/640 [==============================] - 0s 142us/step - loss: 0.5986 - binary_accuracy: 0.7156 - sensitivity: 1.0000 - specificity: 0.0000e+00 - gmeasure: 0.0000e+00 - val_loss: 0.5757 - val_binary_accuracy: 0.7375 - val_sensitivity: 1.0000 - val_specificity: 0.0000e+00 - val_gmeasure: 0.0000e+00
Epoch 4/10
640/640 [==============================] - 0s 127us/step - loss: 0.5984 - binary_accuracy: 0.7156 - sensitivity: 1.0000 - specificity: 0.0000e+00 - gmeasure: 0.0000e+00 - val_loss: 0.5782 - val_binary_accuracy: 0.7375 - val_sensitivity: 1.0000 - val_specificity: 0.0000e+00 - val_gmeasure: 0.0000e+00
Epoch 5/10
640/640 [==============================] - 0s 128us/step - loss: 0.5978 - binary_accuracy: 0.7156 - sensitivity: 1.0000 - specificity: 0.0000e+00 - gmeasure: 0.0000e+00 - val_loss: 0.5766 - val_binary_accuracy: 0.7375 - val_sensitivity: 1.0000 - val_specificity: 0.0000e+00 - val_gmeasure: 0.0000e+00
Epoch 6/10
640/640 [==============================] - 0s 135us/step - loss: 0.5974 - binary_accuracy: 0.7156 - sensitivity: 1.0000 - specificity: 0.0000e+00 - gmeasure: 0.0000e+00 - val_loss: 0.5773 - val_binary_accuracy: 0.7375 - val_sensitivity: 1.0000 - val_specificity: 0.0000e+00 - val_gmeasure: 0.0000e+00
Epoch 7/10
640/640 [==============================] - 0s 138us/step - loss: 0.5976 - binary_accuracy: 0.7156 - sensitivity: 1.0000 - specificity: 0.0000e+00 - gmeasure: 0.0000e+00 - val_loss: 0.5764 - val_binary_accuracy: 0.7375 - val_sensitivity: 1.0000 - val_specificity: 0.0000e+00 - val_gmeasure: 0.0000e+00
Epoch 8/10
640/640 [==============================] - 0s 132us/step - loss: 0.5971 - binary_accuracy: 0.7156 - sensitivity: 1.0000 - specificity: 0.0000e+00 - gmeasure: 0.0000e+00 - val_loss: 0.5770 - val_binary_accuracy: 0.7375 - val_sensitivity: 1.0000 - val_specificity: 0.0000e+00 - val_gmeasure: 0.0000e+00
Epoch 9/10
640/640 [==============================] - 0s 134us/step - loss: 0.5972 - binary_accuracy: 0.7156 - sensitivity: 1.0000 - specificity: 0.0000e+00 - gmeasure: 0.0000e+00 - val_loss: 0.5774 - val_binary_accuracy: 0.7375 - val_sensitivity: 1.0000 - val_specificity: 0.0000e+00 - val_gmeasure: 0.0000e+00
Epoch 10/10
640/640 [==============================] - 0s 145us/step - loss: 0.5975 - binary_accuracy: 0.7156 - sensitivity: 1.0000 - specificity: 0.0000e+00 - gmeasure: 0.0000e+00 - val_loss: 0.5778 - val_binary_accuracy: 0.7375 - val_sensitivity: 1.0000 - val_specificity: 0.0000e+00 - val_gmeasure: 0.0000e+00
[root |INFO|build_network.py:87] Load trained model weight at ./example_result/weight_1.h5
[root |INFO|build_network.py:147] Training end with time 2.7704906463623047!
[root |INFO|build_network.py:83] Saved trained model weight at ./example_result/weight_1.h5
[root |DEBUG|deepbiome.py:185] Save weight at ./example_result/weight_1.h5
[root |DEBUG|deepbiome.py:188] Save history at ./example_result/hist_1.json
[root |INFO|build_network.py:173] Evaluation start!
800/800 [==============================] - 0s 3us/step
[root |INFO|build_network.py:178] Evaluation end with time 0.010298490524291992!
[root |INFO|build_network.py:179] Evaluation: [0.5934057831764221, 0.7200000286102295, 1.0, 0.0, 0.0]
[root |INFO|build_network.py:173] Evaluation start!
200/200 [==============================] - 0s 12us/step
[root |INFO|build_network.py:178] Evaluation end with time 0.009493350982666016!
[root |INFO|build_network.py:179] Evaluation: [0.5934092998504639, 0.7200000286102295, 1.0, 0.0, 0.0]
[root |INFO|deepbiome.py:199] Compute time : 3.284306764602661
[root |INFO|deepbiome.py:200] 2 fold computing end!---------------------------------------------
[root |INFO|deepbiome.py:153] -------3 simulation start!----------------------------------
[root |INFO|readers.py:58] -----------------------------------------------------------------------
[root |INFO|readers.py:59] Construct Dataset
[root |INFO|readers.py:60] -----------------------------------------------------------------------
[root |INFO|readers.py:61] Load data
[root |INFO|deepbiome.py:164] -----------------------------------------------------------------
[root |INFO|deepbiome.py:165] Build network for 3 simulation
[root |INFO|build_network.py:521] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:522] Read phylogenetic tree information from /DATA/home/muha/github_repos/deepbiome/deepbiome/tests/data/genus48_dic.csv
[root |INFO|build_network.py:528] Phylogenetic tree level list: ['Genus', 'Family', 'Order', 'Class', 'Phylum']
[root |INFO|build_network.py:529] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:537] Genus: 48
[root |INFO|build_network.py:537] Family: 40
[root |INFO|build_network.py:537] Order: 23
[root |INFO|build_network.py:537] Class: 17
[root |INFO|build_network.py:537] Phylum: 9
[root |INFO|build_network.py:546] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:547] Phylogenetic_tree_dict info: ['Genus', 'Order', 'Class', 'Family', 'Phylum', 'Number']
[root |INFO|build_network.py:548] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:558] Build edge weights between [ Genus, Family]
[root |INFO|build_network.py:558] Build edge weights between [Family, Order]
[root |INFO|build_network.py:558] Build edge weights between [ Order, Class]
[root |INFO|build_network.py:558] Build edge weights between [ Class, Phylum]
[root |INFO|build_network.py:571] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:586] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:587] Build network based on phylogenetic tree information
[root |INFO|build_network.py:588] ------------------------------------------------------------------------------------------
Model: "model_1"
_________________________________________________________________
Layer (type) Output Shape Param #
[root |INFO|build_network.py:670] ------------------------------------------------------------------------------------------
=================================================================
input (InputLayer) (None, 48) 0
_________________________________________________________________
l1_dense (Dense_with_tree) (None, 40) 1960
_________________________________________________________________
l1_activation (Activation) (None, 40) 0
_________________________________________________________________
l2_dense (Dense_with_tree) (None, 23) 943
_________________________________________________________________
l2_activation (Activation) (None, 23) 0
_________________________________________________________________
l3_dense (Dense_with_tree) (None, 17) 408
_________________________________________________________________
l3_activation (Activation) (None, 17) 0
_________________________________________________________________
l4_dense (Dense_with_tree) (None, 9) 162
_________________________________________________________________
l4_activation (Activation) (None, 9) 0
_________________________________________________________________
last_dense_h (Dense) (None, 1) 10
_________________________________________________________________
p_hat (Activation) (None, 1) 0
=================================================================
Total params: 3,483
Trainable params: 3,483
Non-trainable params: 0
_________________________________________________________________
[root |INFO|build_network.py:61] Build Network
[root |INFO|build_network.py:62] Optimizer = adam
[root |INFO|build_network.py:63] Loss = binary_crossentropy
[root |INFO|build_network.py:64] Metrics = binary_accuracy, sensitivity, specificity, gmeasure
[root |INFO|deepbiome.py:176] -----------------------------------------------------------------
[root |INFO|deepbiome.py:177] 3 fold computing start!----------------------------------
[root |INFO|build_network.py:137] Training start!
Train on 640 samples, validate on 160 samples
Epoch 1/10
640/640 [==============================] - 1s 792us/step - loss: 0.6701 - binary_accuracy: 0.6609 - sensitivity: 0.9231 - specificity: 0.0714 - gmeasure: 0.0000e+00 - val_loss: 0.6407 - val_binary_accuracy: 0.7000 - val_sensitivity: 1.0000 - val_specificity: 0.0000e+00 - val_gmeasure: 0.0000e+00
Epoch 2/10
640/640 [==============================] - 0s 390us/step - loss: 0.6271 - binary_accuracy: 0.6969 - sensitivity: 1.0000 - specificity: 0.0000e+00 - gmeasure: 0.0000e+00 - val_loss: 0.6137 - val_binary_accuracy: 0.7000 - val_sensitivity: 1.0000 - val_specificity: 0.0000e+00 - val_gmeasure: 0.0000e+00
Epoch 3/10
640/640 [==============================] - 0s 398us/step - loss: 0.6143 - binary_accuracy: 0.6969 - sensitivity: 1.0000 - specificity: 0.0000e+00 - gmeasure: 0.0000e+00 - val_loss: 0.6111 - val_binary_accuracy: 0.7000 - val_sensitivity: 1.0000 - val_specificity: 0.0000e+00 - val_gmeasure: 0.0000e+00
Epoch 4/10
640/640 [==============================] - 0s 112us/step - loss: 0.6141 - binary_accuracy: 0.6969 - sensitivity: 1.0000 - specificity: 0.0000e+00 - gmeasure: 0.0000e+00 - val_loss: 0.6111 - val_binary_accuracy: 0.7000 - val_sensitivity: 1.0000 - val_specificity: 0.0000e+00 - val_gmeasure: 0.0000e+00
Epoch 5/10
640/640 [==============================] - 0s 110us/step - loss: 0.6137 - binary_accuracy: 0.6969 - sensitivity: 1.0000 - specificity: 0.0000e+00 - gmeasure: 0.0000e+00 - val_loss: 0.6109 - val_binary_accuracy: 0.7000 - val_sensitivity: 1.0000 - val_specificity: 0.0000e+00 - val_gmeasure: 0.0000e+00
Epoch 6/10
640/640 [==============================] - 0s 140us/step - loss: 0.6137 - binary_accuracy: 0.6969 - sensitivity: 1.0000 - specificity: 0.0000e+00 - gmeasure: 0.0000e+00 - val_loss: 0.6113 - val_binary_accuracy: 0.7000 - val_sensitivity: 1.0000 - val_specificity: 0.0000e+00 - val_gmeasure: 0.0000e+00
Epoch 7/10
640/640 [==============================] - 0s 146us/step - loss: 0.6137 - binary_accuracy: 0.6969 - sensitivity: 1.0000 - specificity: 0.0000e+00 - gmeasure: 0.0000e+00 - val_loss: 0.6110 - val_binary_accuracy: 0.7000 - val_sensitivity: 1.0000 - val_specificity: 0.0000e+00 - val_gmeasure: 0.0000e+00
Epoch 8/10
640/640 [==============================] - 0s 148us/step - loss: 0.6151 - binary_accuracy: 0.6969 - sensitivity: 1.0000 - specificity: 0.0000e+00 - gmeasure: 0.0000e+00 - val_loss: 0.6110 - val_binary_accuracy: 0.7000 - val_sensitivity: 1.0000 - val_specificity: 0.0000e+00 - val_gmeasure: 0.0000e+00
Epoch 9/10
640/640 [==============================] - 0s 108us/step - loss: 0.6147 - binary_accuracy: 0.6969 - sensitivity: 1.0000 - specificity: 0.0000e+00 - gmeasure: 0.0000e+00 - val_loss: 0.6111 - val_binary_accuracy: 0.7000 - val_sensitivity: 1.0000 - val_specificity: 0.0000e+00 - val_gmeasure: 0.0000e+00
Epoch 10/10
640/640 [==============================] - 0s 310us/step - loss: 0.6151 - binary_accuracy: 0.6969 - sensitivity: 1.0000 - specificity: 0.0000e+00 - gmeasure: 0.0000e+00 - val_loss: 0.6109 - val_binary_accuracy: 0.7000 - val_sensitivity: 1.0000 - val_specificity: 0.0000e+00 - val_gmeasure: 0.0000e+00
[root |INFO|build_network.py:87] Load trained model weight at ./example_result/weight_2.h5
[root |INFO|build_network.py:147] Training end with time 3.3226778507232666!
[root |INFO|build_network.py:83] Saved trained model weight at ./example_result/weight_2.h5
[root |DEBUG|deepbiome.py:185] Save weight at ./example_result/weight_2.h5
[root |DEBUG|deepbiome.py:188] Save history at ./example_result/hist_2.json
[root |INFO|build_network.py:173] Evaluation start!
800/800 [==============================] - 0s 9us/step
[root |INFO|build_network.py:178] Evaluation end with time 0.017215967178344727!
[root |INFO|build_network.py:179] Evaluation: [0.6130363345146179, 0.6974999904632568, 1.0, 0.0, 0.0]
[root |INFO|build_network.py:173] Evaluation start!
200/200 [==============================] - 0s 35us/step
[root |INFO|build_network.py:178] Evaluation end with time 0.018899202346801758!
[root |INFO|build_network.py:179] Evaluation: [0.6496703624725342, 0.6549999713897705, 1.0, 0.0, 0.0]
[root |INFO|deepbiome.py:199] Compute time : 4.089082717895508
[root |INFO|deepbiome.py:200] 3 fold computing end!---------------------------------------------
[root |INFO|deepbiome.py:153] -------4 simulation start!----------------------------------
[root |INFO|readers.py:58] -----------------------------------------------------------------------
[root |INFO|readers.py:59] Construct Dataset
[root |INFO|readers.py:60] -----------------------------------------------------------------------
[root |INFO|readers.py:61] Load data
[root |INFO|deepbiome.py:164] -----------------------------------------------------------------
[root |INFO|deepbiome.py:165] Build network for 4 simulation
[root |INFO|build_network.py:521] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:522] Read phylogenetic tree information from /DATA/home/muha/github_repos/deepbiome/deepbiome/tests/data/genus48_dic.csv
[root |INFO|build_network.py:528] Phylogenetic tree level list: ['Genus', 'Family', 'Order', 'Class', 'Phylum']
[root |INFO|build_network.py:529] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:537] Genus: 48
[root |INFO|build_network.py:537] Family: 40
[root |INFO|build_network.py:537] Order: 23
[root |INFO|build_network.py:537] Class: 17
[root |INFO|build_network.py:537] Phylum: 9
[root |INFO|build_network.py:546] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:547] Phylogenetic_tree_dict info: ['Genus', 'Order', 'Class', 'Family', 'Phylum', 'Number']
[root |INFO|build_network.py:548] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:558] Build edge weights between [ Genus, Family]
[root |INFO|build_network.py:558] Build edge weights between [Family, Order]
[root |INFO|build_network.py:558] Build edge weights between [ Order, Class]
[root |INFO|build_network.py:558] Build edge weights between [ Class, Phylum]
[root |INFO|build_network.py:571] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:586] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:587] Build network based on phylogenetic tree information
[root |INFO|build_network.py:588] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:670] ------------------------------------------------------------------------------------------
Model: "model_1"
_________________________________________________________________
Layer (type) Output Shape Param #
=================================================================
input (InputLayer) (None, 48) 0
_________________________________________________________________
l1_dense (Dense_with_tree) (None, 40) 1960
_________________________________________________________________
l1_activation (Activation) (None, 40) 0
_________________________________________________________________
l2_dense (Dense_with_tree) (None, 23) 943
_________________________________________________________________
l2_activation (Activation) (None, 23) 0
_________________________________________________________________
l3_dense (Dense_with_tree) (None, 17) 408
_________________________________________________________________
l3_activation (Activation) (None, 17) 0
_________________________________________________________________
l4_dense (Dense_with_tree) (None, 9) 162
_________________________________________________________________
l4_activation (Activation) (None, 9) 0
_________________________________________________________________
last_dense_h (Dense) (None, 1) 10
_________________________________________________________________
p_hat (Activation) (None, 1) 0
=================================================================
Total params: 3,483
Trainable params: 3,483
Non-trainable params: 0
_________________________________________________________________
[root |INFO|build_network.py:61] Build Network
[root |INFO|build_network.py:62] Optimizer = adam
[root |INFO|build_network.py:63] Loss = binary_crossentropy
[root |INFO|build_network.py:64] Metrics = binary_accuracy, sensitivity, specificity, gmeasure
[root |INFO|deepbiome.py:176] -----------------------------------------------------------------
[root |INFO|deepbiome.py:177] 4 fold computing start!----------------------------------
[root |INFO|build_network.py:137] Training start!
Train on 640 samples, validate on 160 samples
Epoch 1/10
640/640 [==============================] - 1s 1ms/step - loss: 0.6553 - binary_accuracy: 0.6844 - sensitivity: 0.9976 - specificity: 0.0000e+00 - gmeasure: 0.0000e+00 - val_loss: 0.6216 - val_binary_accuracy: 0.6875 - val_sensitivity: 1.0000 - val_specificity: 0.0000e+00 - val_gmeasure: 0.0000e+00
Epoch 2/10
640/640 [==============================] - 0s 330us/step - loss: 0.6256 - binary_accuracy: 0.6859 - sensitivity: 1.0000 - specificity: 0.0000e+00 - gmeasure: 0.0000e+00 - val_loss: 0.6219 - val_binary_accuracy: 0.6875 - val_sensitivity: 1.0000 - val_specificity: 0.0000e+00 - val_gmeasure: 0.0000e+00
Epoch 3/10
640/640 [==============================] - 0s 494us/step - loss: 0.6227 - binary_accuracy: 0.6859 - sensitivity: 1.0000 - specificity: 0.0000e+00 - gmeasure: 0.0000e+00 - val_loss: 0.6209 - val_binary_accuracy: 0.6875 - val_sensitivity: 1.0000 - val_specificity: 0.0000e+00 - val_gmeasure: 0.0000e+00
Epoch 4/10
640/640 [==============================] - 0s 119us/step - loss: 0.6224 - binary_accuracy: 0.6859 - sensitivity: 1.0000 - specificity: 0.0000e+00 - gmeasure: 0.0000e+00 - val_loss: 0.6209 - val_binary_accuracy: 0.6875 - val_sensitivity: 1.0000 - val_specificity: 0.0000e+00 - val_gmeasure: 0.0000e+00
Epoch 5/10
640/640 [==============================] - 0s 259us/step - loss: 0.6223 - binary_accuracy: 0.6859 - sensitivity: 1.0000 - specificity: 0.0000e+00 - gmeasure: 0.0000e+00 - val_loss: 0.6205 - val_binary_accuracy: 0.6875 - val_sensitivity: 1.0000 - val_specificity: 0.0000e+00 - val_gmeasure: 0.0000e+00
Epoch 6/10
640/640 [==============================] - 0s 493us/step - loss: 0.6222 - binary_accuracy: 0.6859 - sensitivity: 1.0000 - specificity: 0.0000e+00 - gmeasure: 0.0000e+00 - val_loss: 0.6203 - val_binary_accuracy: 0.6875 - val_sensitivity: 1.0000 - val_specificity: 0.0000e+00 - val_gmeasure: 0.0000e+00
Epoch 7/10
640/640 [==============================] - 0s 107us/step - loss: 0.6205 - binary_accuracy: 0.6859 - sensitivity: 1.0000 - specificity: 0.0000e+00 - gmeasure: 0.0000e+00 - val_loss: 0.6190 - val_binary_accuracy: 0.6875 - val_sensitivity: 1.0000 - val_specificity: 0.0000e+00 - val_gmeasure: 0.0000e+00
Epoch 8/10
640/640 [==============================] - 0s 103us/step - loss: 0.6185 - binary_accuracy: 0.6859 - sensitivity: 1.0000 - specificity: 0.0000e+00 - gmeasure: 0.0000e+00 - val_loss: 0.6155 - val_binary_accuracy: 0.6875 - val_sensitivity: 1.0000 - val_specificity: 0.0000e+00 - val_gmeasure: 0.0000e+00
Epoch 9/10
640/640 [==============================] - 0s 393us/step - loss: 0.6139 - binary_accuracy: 0.6859 - sensitivity: 1.0000 - specificity: 0.0000e+00 - gmeasure: 0.0000e+00 - val_loss: 0.6095 - val_binary_accuracy: 0.6875 - val_sensitivity: 1.0000 - val_specificity: 0.0000e+00 - val_gmeasure: 0.0000e+00
Epoch 10/10
640/640 [==============================] - 0s 372us/step - loss: 0.6019 - binary_accuracy: 0.6859 - sensitivity: 1.0000 - specificity: 0.0000e+00 - gmeasure: 0.0000e+00 - val_loss: 0.5981 - val_binary_accuracy: 0.6875 - val_sensitivity: 1.0000 - val_specificity: 0.0000e+00 - val_gmeasure: 0.0000e+00
[root |INFO|build_network.py:87] Load trained model weight at ./example_result/weight_3.h5
[root |INFO|build_network.py:147] Training end with time 4.37174129486084!
[root |INFO|build_network.py:83] Saved trained model weight at ./example_result/weight_3.h5
[root |DEBUG|deepbiome.py:185] Save weight at ./example_result/weight_3.h5
[root |DEBUG|deepbiome.py:188] Save history at ./example_result/hist_3.json
[root |INFO|build_network.py:173] Evaluation start!
800/800 [==============================] - 0s 19us/step
[root |INFO|build_network.py:178] Evaluation end with time 0.026946306228637695!
[root |INFO|build_network.py:179] Evaluation: [0.5943012833595276, 0.6862499713897705, 1.0, 0.0, 0.0]
[root |INFO|build_network.py:173] Evaluation start!
200/200 [==============================] - 0s 13us/step
[root |INFO|build_network.py:178] Evaluation end with time 0.010583877563476562!
[root |INFO|build_network.py:179] Evaluation: [0.5873210430145264, 0.6899999976158142, 1.0, 0.0, 0.0]
[root |INFO|deepbiome.py:199] Compute time : 4.833027362823486
[root |INFO|deepbiome.py:200] 4 fold computing end!---------------------------------------------
[root |INFO|deepbiome.py:153] -------5 simulation start!----------------------------------
[root |INFO|readers.py:58] -----------------------------------------------------------------------
[root |INFO|readers.py:59] Construct Dataset
[root |INFO|readers.py:60] -----------------------------------------------------------------------
[root |INFO|readers.py:61] Load data
[root |INFO|deepbiome.py:164] -----------------------------------------------------------------
[root |INFO|deepbiome.py:165] Build network for 5 simulation
[root |INFO|build_network.py:521] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:522] Read phylogenetic tree information from /DATA/home/muha/github_repos/deepbiome/deepbiome/tests/data/genus48_dic.csv
[root |INFO|build_network.py:528] Phylogenetic tree level list: ['Genus', 'Family', 'Order', 'Class', 'Phylum']
[root |INFO|build_network.py:529] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:537] Genus: 48
[root |INFO|build_network.py:537] Family: 40
[root |INFO|build_network.py:537] Order: 23
[root |INFO|build_network.py:537] Class: 17
[root |INFO|build_network.py:537] Phylum: 9
[root |INFO|build_network.py:546] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:547] Phylogenetic_tree_dict info: ['Genus', 'Order', 'Class', 'Family', 'Phylum', 'Number']
[root |INFO|build_network.py:548] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:558] Build edge weights between [ Genus, Family]
[root |INFO|build_network.py:558] Build edge weights between [Family, Order]
[root |INFO|build_network.py:558] Build edge weights between [ Order, Class]
[root |INFO|build_network.py:558] Build edge weights between [ Class, Phylum]
[root |INFO|build_network.py:571] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:586] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:587] Build network based on phylogenetic tree information
[root |INFO|build_network.py:588] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:670] ------------------------------------------------------------------------------------------
Model: "model_1"
_________________________________________________________________
Layer (type) Output Shape Param #
=================================================================
input (InputLayer) (None, 48) 0
_________________________________________________________________
l1_dense (Dense_with_tree) (None, 40) 1960
_________________________________________________________________
l1_activation (Activation) (None, 40) 0
_________________________________________________________________
l2_dense (Dense_with_tree) (None, 23) 943
_________________________________________________________________
l2_activation (Activation) (None, 23) 0
_________________________________________________________________
l3_dense (Dense_with_tree) (None, 17) 408
_________________________________________________________________
l3_activation (Activation) (None, 17) 0
_________________________________________________________________
l4_dense (Dense_with_tree) (None, 9) 162
_________________________________________________________________
l4_activation (Activation) (None, 9) 0
_________________________________________________________________
last_dense_h (Dense) (None, 1) 10
_________________________________________________________________
p_hat (Activation) (None, 1) 0
=================================================================
Total params: 3,483
Trainable params: 3,483
Non-trainable params: 0
_________________________________________________________________
[root |INFO|build_network.py:61] Build Network
[root |INFO|build_network.py:62] Optimizer = adam
[root |INFO|build_network.py:63] Loss = binary_crossentropy
[root |INFO|build_network.py:64] Metrics = binary_accuracy, sensitivity, specificity, gmeasure
[root |INFO|deepbiome.py:176] -----------------------------------------------------------------
[root |INFO|deepbiome.py:177] 5 fold computing start!----------------------------------
[root |INFO|build_network.py:137] Training start!
Train on 640 samples, validate on 160 samples
Epoch 1/10
640/640 [==============================] - 1s 1ms/step - loss: 0.6673 - binary_accuracy: 0.6797 - sensitivity: 1.0000 - specificity: 0.0000e+00 - gmeasure: 0.0000e+00 - val_loss: 0.6302 - val_binary_accuracy: 0.6938 - val_sensitivity: 1.0000 - val_specificity: 0.0000e+00 - val_gmeasure: 0.0000e+00
Epoch 2/10
640/640 [==============================] - 0s 111us/step - loss: 0.6310 - binary_accuracy: 0.6797 - sensitivity: 1.0000 - specificity: 0.0000e+00 - gmeasure: 0.0000e+00 - val_loss: 0.6165 - val_binary_accuracy: 0.6938 - val_sensitivity: 1.0000 - val_specificity: 0.0000e+00 - val_gmeasure: 0.0000e+00
Epoch 3/10
640/640 [==============================] - 0s 106us/step - loss: 0.6284 - binary_accuracy: 0.6797 - sensitivity: 1.0000 - specificity: 0.0000e+00 - gmeasure: 0.0000e+00 - val_loss: 0.6163 - val_binary_accuracy: 0.6938 - val_sensitivity: 1.0000 - val_specificity: 0.0000e+00 - val_gmeasure: 0.0000e+00
Epoch 4/10
640/640 [==============================] - 0s 129us/step - loss: 0.6275 - binary_accuracy: 0.6797 - sensitivity: 1.0000 - specificity: 0.0000e+00 - gmeasure: 0.0000e+00 - val_loss: 0.6173 - val_binary_accuracy: 0.6938 - val_sensitivity: 1.0000 - val_specificity: 0.0000e+00 - val_gmeasure: 0.0000e+00
Epoch 5/10
640/640 [==============================] - 0s 145us/step - loss: 0.6274 - binary_accuracy: 0.6797 - sensitivity: 1.0000 - specificity: 0.0000e+00 - gmeasure: 0.0000e+00 - val_loss: 0.6167 - val_binary_accuracy: 0.6938 - val_sensitivity: 1.0000 - val_specificity: 0.0000e+00 - val_gmeasure: 0.0000e+00
Epoch 6/10
640/640 [==============================] - 0s 156us/step - loss: 0.6276 - binary_accuracy: 0.6797 - sensitivity: 1.0000 - specificity: 0.0000e+00 - gmeasure: 0.0000e+00 - val_loss: 0.6168 - val_binary_accuracy: 0.6938 - val_sensitivity: 1.0000 - val_specificity: 0.0000e+00 - val_gmeasure: 0.0000e+00
Epoch 7/10
640/640 [==============================] - 0s 163us/step - loss: 0.6272 - binary_accuracy: 0.6797 - sensitivity: 1.0000 - specificity: 0.0000e+00 - gmeasure: 0.0000e+00 - val_loss: 0.6163 - val_binary_accuracy: 0.6938 - val_sensitivity: 1.0000 - val_specificity: 0.0000e+00 - val_gmeasure: 0.0000e+00
Epoch 8/10
640/640 [==============================] - 0s 170us/step - loss: 0.6274 - binary_accuracy: 0.6797 - sensitivity: 1.0000 - specificity: 0.0000e+00 - gmeasure: 0.0000e+00 - val_loss: 0.6162 - val_binary_accuracy: 0.6938 - val_sensitivity: 1.0000 - val_specificity: 0.0000e+00 - val_gmeasure: 0.0000e+00
Epoch 9/10
640/640 [==============================] - 0s 141us/step - loss: 0.6273 - binary_accuracy: 0.6797 - sensitivity: 1.0000 - specificity: 0.0000e+00 - gmeasure: 0.0000e+00 - val_loss: 0.6165 - val_binary_accuracy: 0.6938 - val_sensitivity: 1.0000 - val_specificity: 0.0000e+00 - val_gmeasure: 0.0000e+00
Epoch 10/10
640/640 [==============================] - 0s 132us/step - loss: 0.6279 - binary_accuracy: 0.6797 - sensitivity: 1.0000 - specificity: 0.0000e+00 - gmeasure: 0.0000e+00 - val_loss: 0.6163 - val_binary_accuracy: 0.6938 - val_sensitivity: 1.0000 - val_specificity: 0.0000e+00 - val_gmeasure: 0.0000e+00
[root |INFO|build_network.py:87] Load trained model weight at ./example_result/weight_4.h5
[root |INFO|build_network.py:147] Training end with time 3.268160104751587!
[root |INFO|build_network.py:83] Saved trained model weight at ./example_result/weight_4.h5
[root |DEBUG|deepbiome.py:185] Save weight at ./example_result/weight_4.h5
[root |DEBUG|deepbiome.py:188] Save history at ./example_result/hist_4.json
[root |INFO|build_network.py:173] Evaluation start!
800/800 [==============================] - 0s 5us/step
[root |INFO|build_network.py:178] Evaluation end with time 0.011313438415527344!
[root |INFO|build_network.py:179] Evaluation: [0.6249020099639893, 0.6825000047683716, 1.0, 0.0, 0.0]
[root |INFO|build_network.py:173] Evaluation start!
200/200 [==============================] - 0s 15us/step
[root |INFO|build_network.py:178] Evaluation end with time 0.010974407196044922!
[root |INFO|build_network.py:179] Evaluation: [0.6075432896614075, 0.7049999833106995, 1.0, 0.0, 0.0]
[root |INFO|deepbiome.py:199] Compute time : 3.72648286819458
[root |INFO|deepbiome.py:200] 5 fold computing end!---------------------------------------------
[root |INFO|deepbiome.py:211] -----------------------------------------------------------------
[root |INFO|deepbiome.py:212] Train Evaluation : ['loss' 'binary_accuracy' 'sensitivity' 'specificity' 'gmeasure']
[root |INFO|deepbiome.py:213] mean : [0.609 0.694 1.000 0.000 0.000]
[root |INFO|deepbiome.py:214] std : [0.013 0.014 0.000 0.000 0.000]
[root |INFO|deepbiome.py:215] -----------------------------------------------------------------
[root |INFO|deepbiome.py:216] Test Evaluation : ['loss' 'binary_accuracy' 'sensitivity' 'specificity' 'gmeasure']
[root |INFO|deepbiome.py:217] mean : [0.611 0.692 1.000 0.000 0.000]
[root |INFO|deepbiome.py:218] std : [0.022 0.022 0.000 0.000 0.000]
[root |INFO|deepbiome.py:219] -----------------------------------------------------------------
[root |INFO|deepbiome.py:230] Total Computing Ended
[root |INFO|deepbiome.py:231] -----------------------------------------------------------------
The deepbiome_train saves the trained model weights, evaluation results and history based on the path information from the configuration.
From the example above, we can check that hist_*.json, weight_*.h5, test_eval.npy, train_eval.npy files were saved.
[19]:
os.listdir(path_info['model_info']['model_dir'])
[19]:
['hist_0.json',
'weight_2.h5',
'test_eval.npy',
'weight_0.h5',
'train_eval.npy',
'hist_2.json',
'hist_4.json',
'weight_3.h5',
'hist_3.json',
'weight_1.h5',
'weight_4.h5',
'hist_1.json']
Lets check the history files.
[20]:
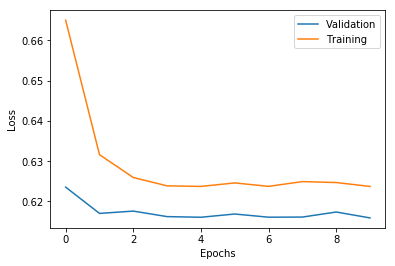
with open('./%s/hist_0.json' % path_info['model_info']['model_dir'], 'r') as f:
history = json.load(f)
plt.plot(history['val_loss'], label='Validation')
plt.plot(history['loss'], label='Training')
plt.legend()
plt.xlabel('Epochs')
plt.ylabel('Loss')
plt.show()
Test evauation and train evauation is the numpy array of the shape (number of folds, number of evaluation measures).
[21]:
test_evaluation
[21]:
array([[0.619, 0.690, 1.000, 0.000, 0.000],
[0.593, 0.720, 1.000, 0.000, 0.000],
[0.650, 0.655, 1.000, 0.000, 0.000],
[0.587, 0.690, 1.000, 0.000, 0.000],
[0.608, 0.705, 1.000, 0.000, 0.000]])
[22]:
train_evaluation
[22]:
array([[0.622, 0.686, 1.000, 0.000, 0.000],
[0.593, 0.720, 1.000, 0.000, 0.000],
[0.613, 0.697, 1.000, 0.000, 0.000],
[0.594, 0.686, 1.000, 0.000, 0.000],
[0.625, 0.683, 1.000, 0.000, 0.000]])
5. Load the pre-trained network for training¶
If you have a pre-trianed model, you warm_start next training using the pre-trained weights by setting the warm_start option in training_info to True. The file path of the pre-trained weights passed in the warm_start_model option. Below is the example:
[23]:
warm_start_network_info = {
'architecture_info': {
'batch_normalization': 'False',
'drop_out': '0',
'weight_initial': 'glorot_uniform',
'weight_l1_penalty':'0.',
'weight_decay': 'phylogenetic_tree',
},
'model_info': {
'decay': '0.001',
'loss': 'binary_crossentropy',
'lr': '0.01',
'metrics': 'binary_accuracy, sensitivity, specificity, gmeasure',
'network_class': 'DeepBiomeNetwork',
'normalizer': 'normalize_minmax',
'optimizer': 'adam',
'reader_class': 'MicroBiomeClassificationReader',
'taxa_selection_metrics': 'sensitivity, specificity, gmeasure, accuracy',
},
'training_info': {
'warm_start':'True',
'warm_start_model':'./example_result/weight.h5',
'epochs': '10',
'batch_size': '50',
'callbacks': 'ModelCheckpoint',
'monitor': 'val_loss',
'mode' : 'min',
'min_delta': '1e-7',
},
'validation_info': {
'batch_size': 'None',
'validation_size': '0.2'
},
'test_info': {
'batch_size': 'None'
}
}
[24]:
test_evaluation, train_evaluation, network = deepbiome.deepbiome_train(log, warm_start_network_info, path_info,
number_of_fold=None)
[root |INFO|deepbiome.py:115] -----------------------------------------------------------------
[root |INFO|deepbiome.py:153] -------1 simulation start!----------------------------------
[root |INFO|readers.py:58] -----------------------------------------------------------------------
[root |INFO|readers.py:59] Construct Dataset
[root |INFO|readers.py:60] -----------------------------------------------------------------------
[root |INFO|readers.py:61] Load data
[root |INFO|deepbiome.py:164] -----------------------------------------------------------------
[root |INFO|deepbiome.py:165] Build network for 1 simulation
[root |INFO|build_network.py:521] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:522] Read phylogenetic tree information from /DATA/home/muha/github_repos/deepbiome/deepbiome/tests/data/genus48_dic.csv
[root |INFO|build_network.py:528] Phylogenetic tree level list: ['Genus', 'Family', 'Order', 'Class', 'Phylum']
[root |INFO|build_network.py:529] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:537] Genus: 48
[root |INFO|build_network.py:537] Family: 40
[root |INFO|build_network.py:537] Order: 23
[root |INFO|build_network.py:537] Class: 17
[root |INFO|build_network.py:537] Phylum: 9
[root |INFO|build_network.py:546] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:547] Phylogenetic_tree_dict info: ['Genus', 'Order', 'Class', 'Family', 'Phylum', 'Number']
[root |INFO|build_network.py:548] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:558] Build edge weights between [ Genus, Family]
[root |INFO|build_network.py:558] Build edge weights between [Family, Order]
[root |INFO|build_network.py:558] Build edge weights between [ Order, Class]
[root |INFO|build_network.py:558] Build edge weights between [ Class, Phylum]
[root |INFO|build_network.py:571] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:586] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:587] Build network based on phylogenetic tree information
[root |INFO|build_network.py:588] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:670] ------------------------------------------------------------------------------------------
Model: "model_1"
_________________________________________________________________
Layer (type) Output Shape Param #
=================================================================
input (InputLayer) (None, 48) 0
_________________________________________________________________
l1_dense (Dense_with_tree) (None, 40) 1960
_________________________________________________________________
l1_activation (Activation) (None, 40) 0
_________________________________________________________________
l2_dense (Dense_with_tree) (None, 23) 943
_________________________________________________________________
l2_activation (Activation) (None, 23) 0
_________________________________________________________________
l3_dense (Dense_with_tree) (None, 17) 408
_________________________________________________________________
l3_activation (Activation) (None, 17) 0
_________________________________________________________________
l4_dense (Dense_with_tree) (None, 9) 162
_________________________________________________________________
l4_activation (Activation) (None, 9) 0
_________________________________________________________________
last_dense_h (Dense) (None, 1) 10
_________________________________________________________________
p_hat (Activation) (None, 1) 0
=================================================================
Total params: 3,483
Trainable params: 3,483
Non-trainable params: 0
_________________________________________________________________
[root |INFO|build_network.py:61] Build Network
[root |INFO|build_network.py:62] Optimizer = adam
[root |INFO|build_network.py:63] Loss = binary_crossentropy
[root |INFO|build_network.py:64] Metrics = binary_accuracy, sensitivity, specificity, gmeasure
[root |INFO|build_network.py:87] Load trained model weight at ./example_result/weight_0.h5
[root |INFO|deepbiome.py:176] -----------------------------------------------------------------
[root |INFO|deepbiome.py:177] 1 fold computing start!----------------------------------
[root |INFO|build_network.py:137] Training start!
Train on 640 samples, validate on 160 samples
Epoch 1/10
640/640 [==============================] - 0s 769us/step - loss: 0.6264 - binary_accuracy: 0.6844 - sensitivity: 1.0000 - specificity: 0.0000e+00 - gmeasure: 0.0000e+00 - val_loss: 0.6162 - val_binary_accuracy: 0.6938 - val_sensitivity: 1.0000 - val_specificity: 0.0000e+00 - val_gmeasure: 0.0000e+00
Epoch 2/10
640/640 [==============================] - 0s 138us/step - loss: 0.6246 - binary_accuracy: 0.6844 - sensitivity: 1.0000 - specificity: 0.0000e+00 - gmeasure: 0.0000e+00 - val_loss: 0.6156 - val_binary_accuracy: 0.6938 - val_sensitivity: 1.0000 - val_specificity: 0.0000e+00 - val_gmeasure: 0.0000e+00
Epoch 3/10
640/640 [==============================] - 0s 152us/step - loss: 0.6226 - binary_accuracy: 0.6844 - sensitivity: 1.0000 - specificity: 0.0000e+00 - gmeasure: 0.0000e+00 - val_loss: 0.6167 - val_binary_accuracy: 0.6938 - val_sensitivity: 1.0000 - val_specificity: 0.0000e+00 - val_gmeasure: 0.0000e+00
Epoch 4/10
640/640 [==============================] - 0s 144us/step - loss: 0.6238 - binary_accuracy: 0.6844 - sensitivity: 1.0000 - specificity: 0.0000e+00 - gmeasure: 0.0000e+00 - val_loss: 0.6160 - val_binary_accuracy: 0.6938 - val_sensitivity: 1.0000 - val_specificity: 0.0000e+00 - val_gmeasure: 0.0000e+00
Epoch 5/10
640/640 [==============================] - 0s 153us/step - loss: 0.6271 - binary_accuracy: 0.6844 - sensitivity: 1.0000 - specificity: 0.0000e+00 - gmeasure: 0.0000e+00 - val_loss: 0.6140 - val_binary_accuracy: 0.6938 - val_sensitivity: 1.0000 - val_specificity: 0.0000e+00 - val_gmeasure: 0.0000e+00
Epoch 6/10
640/640 [==============================] - 0s 159us/step - loss: 0.6199 - binary_accuracy: 0.6844 - sensitivity: 1.0000 - specificity: 0.0000e+00 - gmeasure: 0.0000e+00 - val_loss: 0.6135 - val_binary_accuracy: 0.6938 - val_sensitivity: 1.0000 - val_specificity: 0.0000e+00 - val_gmeasure: 0.0000e+00
Epoch 7/10
640/640 [==============================] - 0s 131us/step - loss: 0.6213 - binary_accuracy: 0.6844 - sensitivity: 1.0000 - specificity: 0.0000e+00 - gmeasure: 0.0000e+00 - val_loss: 0.6118 - val_binary_accuracy: 0.6938 - val_sensitivity: 1.0000 - val_specificity: 0.0000e+00 - val_gmeasure: 0.0000e+00
Epoch 8/10
640/640 [==============================] - 0s 136us/step - loss: 0.6186 - binary_accuracy: 0.6844 - sensitivity: 1.0000 - specificity: 0.0000e+00 - gmeasure: 0.0000e+00 - val_loss: 0.6086 - val_binary_accuracy: 0.6938 - val_sensitivity: 1.0000 - val_specificity: 0.0000e+00 - val_gmeasure: 0.0000e+00
Epoch 9/10
640/640 [==============================] - 0s 143us/step - loss: 0.6157 - binary_accuracy: 0.6844 - sensitivity: 1.0000 - specificity: 0.0000e+00 - gmeasure: 0.0000e+00 - val_loss: 0.6055 - val_binary_accuracy: 0.6938 - val_sensitivity: 1.0000 - val_specificity: 0.0000e+00 - val_gmeasure: 0.0000e+00
Epoch 10/10
640/640 [==============================] - 0s 154us/step - loss: 0.6128 - binary_accuracy: 0.6844 - sensitivity: 1.0000 - specificity: 0.0000e+00 - gmeasure: 0.0000e+00 - val_loss: 0.6016 - val_binary_accuracy: 0.6938 - val_sensitivity: 1.0000 - val_specificity: 0.0000e+00 - val_gmeasure: 0.0000e+00
[root |INFO|build_network.py:87] Load trained model weight at ./example_result/weight_0.h5
[root |INFO|build_network.py:147] Training end with time 2.9706335067749023!
[root |INFO|build_network.py:83] Saved trained model weight at ./example_result/weight_0.h5
[root |DEBUG|deepbiome.py:185] Save weight at ./example_result/weight_0.h5
[root |DEBUG|deepbiome.py:188] Save history at ./example_result/hist_0.json
[root |INFO|build_network.py:173] Evaluation start!
800/800 [==============================] - 0s 4us/step
[root |INFO|build_network.py:178] Evaluation end with time 0.01216745376586914!
[root |INFO|build_network.py:179] Evaluation: [0.6077973246574402, 0.6862499713897705, 1.0, 0.0, 0.0]
[root |INFO|build_network.py:173] Evaluation start!
200/200 [==============================] - 0s 14us/step
[root |INFO|build_network.py:178] Evaluation end with time 0.010446786880493164!
[root |INFO|build_network.py:179] Evaluation: [0.608977735042572, 0.6899999976158142, 1.0, 0.0, 0.0]
[root |INFO|deepbiome.py:199] Compute time : 3.626851797103882
[root |INFO|deepbiome.py:200] 1 fold computing end!---------------------------------------------
[root |INFO|deepbiome.py:153] -------2 simulation start!----------------------------------
[root |INFO|readers.py:58] -----------------------------------------------------------------------
[root |INFO|readers.py:59] Construct Dataset
[root |INFO|readers.py:60] -----------------------------------------------------------------------
[root |INFO|readers.py:61] Load data
[root |INFO|deepbiome.py:164] -----------------------------------------------------------------
[root |INFO|deepbiome.py:165] Build network for 2 simulation
[root |INFO|build_network.py:521] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:522] Read phylogenetic tree information from /DATA/home/muha/github_repos/deepbiome/deepbiome/tests/data/genus48_dic.csv
[root |INFO|build_network.py:528] Phylogenetic tree level list: ['Genus', 'Family', 'Order', 'Class', 'Phylum']
[root |INFO|build_network.py:529] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:537] Genus: 48
[root |INFO|build_network.py:537] Family: 40
[root |INFO|build_network.py:537] Order: 23
[root |INFO|build_network.py:537] Class: 17
[root |INFO|build_network.py:537] Phylum: 9
[root |INFO|build_network.py:546] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:547] Phylogenetic_tree_dict info: ['Genus', 'Order', 'Class', 'Family', 'Phylum', 'Number']
[root |INFO|build_network.py:548] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:558] Build edge weights between [ Genus, Family]
[root |INFO|build_network.py:558] Build edge weights between [Family, Order]
[root |INFO|build_network.py:558] Build edge weights between [ Order, Class]
[root |INFO|build_network.py:558] Build edge weights between [ Class, Phylum]
[root |INFO|build_network.py:571] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:586] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:587] Build network based on phylogenetic tree information
[root |INFO|build_network.py:588] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:670] ------------------------------------------------------------------------------------------
Model: "model_1"
_________________________________________________________________
Layer (type) Output Shape Param #
=================================================================
input (InputLayer) (None, 48) 0
_________________________________________________________________
l1_dense (Dense_with_tree) (None, 40) 1960
_________________________________________________________________
l1_activation (Activation) (None, 40) 0
_________________________________________________________________
l2_dense (Dense_with_tree) (None, 23) 943
_________________________________________________________________
l2_activation (Activation) (None, 23) 0
_________________________________________________________________
l3_dense (Dense_with_tree) (None, 17) 408
_________________________________________________________________
l3_activation (Activation) (None, 17) 0
_________________________________________________________________
l4_dense (Dense_with_tree) (None, 9) 162
_________________________________________________________________
l4_activation (Activation) (None, 9) 0
_________________________________________________________________
last_dense_h (Dense) (None, 1) 10
_________________________________________________________________
p_hat (Activation) (None, 1) 0
=================================================================
Total params: 3,483
Trainable params: 3,483
Non-trainable params: 0
_________________________________________________________________
[root |INFO|build_network.py:61] Build Network
[root |INFO|build_network.py:62] Optimizer = adam
[root |INFO|build_network.py:63] Loss = binary_crossentropy
[root |INFO|build_network.py:64] Metrics = binary_accuracy, sensitivity, specificity, gmeasure
[root |INFO|build_network.py:87] Load trained model weight at ./example_result/weight_1.h5
[root |INFO|deepbiome.py:176] -----------------------------------------------------------------
[root |INFO|deepbiome.py:177] 2 fold computing start!----------------------------------
[root |INFO|build_network.py:137] Training start!
Train on 640 samples, validate on 160 samples
Epoch 1/10
640/640 [==============================] - 1s 856us/step - loss: 0.5983 - binary_accuracy: 0.7156 - sensitivity: 1.0000 - specificity: 0.0000e+00 - gmeasure: 0.0000e+00 - val_loss: 0.5809 - val_binary_accuracy: 0.7375 - val_sensitivity: 1.0000 - val_specificity: 0.0000e+00 - val_gmeasure: 0.0000e+00
Epoch 2/10
640/640 [==============================] - 0s 135us/step - loss: 0.5976 - binary_accuracy: 0.7156 - sensitivity: 1.0000 - specificity: 0.0000e+00 - gmeasure: 0.0000e+00 - val_loss: 0.5766 - val_binary_accuracy: 0.7375 - val_sensitivity: 1.0000 - val_specificity: 0.0000e+00 - val_gmeasure: 0.0000e+00
Epoch 3/10
640/640 [==============================] - 0s 151us/step - loss: 0.5978 - binary_accuracy: 0.7156 - sensitivity: 1.0000 - specificity: 0.0000e+00 - gmeasure: 0.0000e+00 - val_loss: 0.5759 - val_binary_accuracy: 0.7375 - val_sensitivity: 1.0000 - val_specificity: 0.0000e+00 - val_gmeasure: 0.0000e+00
Epoch 4/10
640/640 [==============================] - 0s 135us/step - loss: 0.5980 - binary_accuracy: 0.7156 - sensitivity: 1.0000 - specificity: 0.0000e+00 - gmeasure: 0.0000e+00 - val_loss: 0.5787 - val_binary_accuracy: 0.7375 - val_sensitivity: 1.0000 - val_specificity: 0.0000e+00 - val_gmeasure: 0.0000e+00
Epoch 5/10
640/640 [==============================] - 0s 139us/step - loss: 0.5975 - binary_accuracy: 0.7156 - sensitivity: 1.0000 - specificity: 0.0000e+00 - gmeasure: 0.0000e+00 - val_loss: 0.5767 - val_binary_accuracy: 0.7375 - val_sensitivity: 1.0000 - val_specificity: 0.0000e+00 - val_gmeasure: 0.0000e+00
Epoch 6/10
640/640 [==============================] - 0s 123us/step - loss: 0.5977 - binary_accuracy: 0.7156 - sensitivity: 1.0000 - specificity: 0.0000e+00 - gmeasure: 0.0000e+00 - val_loss: 0.5761 - val_binary_accuracy: 0.7375 - val_sensitivity: 1.0000 - val_specificity: 0.0000e+00 - val_gmeasure: 0.0000e+00
Epoch 7/10
640/640 [==============================] - 0s 108us/step - loss: 0.5986 - binary_accuracy: 0.7156 - sensitivity: 1.0000 - specificity: 0.0000e+00 - gmeasure: 0.0000e+00 - val_loss: 0.5784 - val_binary_accuracy: 0.7375 - val_sensitivity: 1.0000 - val_specificity: 0.0000e+00 - val_gmeasure: 0.0000e+00
Epoch 8/10
640/640 [==============================] - 0s 114us/step - loss: 0.5970 - binary_accuracy: 0.7156 - sensitivity: 1.0000 - specificity: 0.0000e+00 - gmeasure: 0.0000e+00 - val_loss: 0.5763 - val_binary_accuracy: 0.7375 - val_sensitivity: 1.0000 - val_specificity: 0.0000e+00 - val_gmeasure: 0.0000e+00
Epoch 9/10
640/640 [==============================] - 0s 106us/step - loss: 0.5976 - binary_accuracy: 0.7156 - sensitivity: 1.0000 - specificity: 0.0000e+00 - gmeasure: 0.0000e+00 - val_loss: 0.5771 - val_binary_accuracy: 0.7375 - val_sensitivity: 1.0000 - val_specificity: 0.0000e+00 - val_gmeasure: 0.0000e+00
Epoch 10/10
640/640 [==============================] - 0s 127us/step - loss: 0.5973 - binary_accuracy: 0.7156 - sensitivity: 1.0000 - specificity: 0.0000e+00 - gmeasure: 0.0000e+00 - val_loss: 0.5763 - val_binary_accuracy: 0.7375 - val_sensitivity: 1.0000 - val_specificity: 0.0000e+00 - val_gmeasure: 0.0000e+00
[root |INFO|build_network.py:87] Load trained model weight at ./example_result/weight_1.h5
[root |INFO|build_network.py:147] Training end with time 2.6066670417785645!
[root |INFO|build_network.py:83] Saved trained model weight at ./example_result/weight_1.h5
[root |DEBUG|deepbiome.py:185] Save weight at ./example_result/weight_1.h5
[root |DEBUG|deepbiome.py:188] Save history at ./example_result/hist_1.json
[root |INFO|build_network.py:173] Evaluation start!
800/800 [==============================] - 0s 3us/step
[root |INFO|build_network.py:178] Evaluation end with time 0.008038043975830078!
[root |INFO|build_network.py:179] Evaluation: [0.5931535363197327, 0.7200000286102295, 1.0, 0.0, 0.0]
[root |INFO|build_network.py:173] Evaluation start!
200/200 [==============================] - 0s 8us/step
[root |INFO|build_network.py:178] Evaluation end with time 0.009198188781738281!
[root |INFO|build_network.py:179] Evaluation: [0.5931748747825623, 0.7200000286102295, 1.0, 0.0, 0.0]
[root |INFO|deepbiome.py:199] Compute time : 3.5099010467529297
[root |INFO|deepbiome.py:200] 2 fold computing end!---------------------------------------------
[root |INFO|deepbiome.py:153] -------3 simulation start!----------------------------------
[root |INFO|readers.py:58] -----------------------------------------------------------------------
[root |INFO|readers.py:59] Construct Dataset
[root |INFO|readers.py:60] -----------------------------------------------------------------------
[root |INFO|readers.py:61] Load data
[root |INFO|deepbiome.py:164] -----------------------------------------------------------------
[root |INFO|deepbiome.py:165] Build network for 3 simulation
[root |INFO|build_network.py:521] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:522] Read phylogenetic tree information from /DATA/home/muha/github_repos/deepbiome/deepbiome/tests/data/genus48_dic.csv
[root |INFO|build_network.py:528] Phylogenetic tree level list: ['Genus', 'Family', 'Order', 'Class', 'Phylum']
[root |INFO|build_network.py:529] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:537] Genus: 48
[root |INFO|build_network.py:537] Family: 40
[root |INFO|build_network.py:537] Order: 23
[root |INFO|build_network.py:537] Class: 17
[root |INFO|build_network.py:537] Phylum: 9
[root |INFO|build_network.py:546] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:547] Phylogenetic_tree_dict info: ['Genus', 'Order', 'Class', 'Family', 'Phylum', 'Number']
[root |INFO|build_network.py:548] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:558] Build edge weights between [ Genus, Family]
[root |INFO|build_network.py:558] Build edge weights between [Family, Order]
[root |INFO|build_network.py:558] Build edge weights between [ Order, Class]
[root |INFO|build_network.py:558] Build edge weights between [ Class, Phylum]
[root |INFO|build_network.py:571] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:586] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:587] Build network based on phylogenetic tree information
[root |INFO|build_network.py:588] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:670] ------------------------------------------------------------------------------------------
Model: "model_1"
_________________________________________________________________
Layer (type) Output Shape Param #
=================================================================
input (InputLayer) (None, 48) 0
_________________________________________________________________
l1_dense (Dense_with_tree) (None, 40) 1960
_________________________________________________________________
l1_activation (Activation) (None, 40) 0
_________________________________________________________________
l2_dense (Dense_with_tree) (None, 23) 943
_________________________________________________________________
l2_activation (Activation) (None, 23) 0
_________________________________________________________________
l3_dense (Dense_with_tree) (None, 17) 408
_________________________________________________________________
l3_activation (Activation) (None, 17) 0
_________________________________________________________________
l4_dense (Dense_with_tree) (None, 9) 162
_________________________________________________________________
l4_activation (Activation) (None, 9) 0
_________________________________________________________________
last_dense_h (Dense) (None, 1) 10
_________________________________________________________________
p_hat (Activation) (None, 1) 0
=================================================================
Total params: 3,483
Trainable params: 3,483
Non-trainable params: 0
_________________________________________________________________
[root |INFO|build_network.py:61] Build Network
[root |INFO|build_network.py:62] Optimizer = adam
[root |INFO|build_network.py:63] Loss = binary_crossentropy
[root |INFO|build_network.py:64] Metrics = binary_accuracy, sensitivity, specificity, gmeasure
[root |INFO|build_network.py:87] Load trained model weight at ./example_result/weight_2.h5
[root |INFO|deepbiome.py:176] -----------------------------------------------------------------
[root |INFO|deepbiome.py:177] 3 fold computing start!----------------------------------
[root |INFO|build_network.py:137] Training start!
Train on 640 samples, validate on 160 samples
Epoch 1/10
640/640 [==============================] - 1s 841us/step - loss: 0.6138 - binary_accuracy: 0.6969 - sensitivity: 1.0000 - specificity: 0.0000e+00 - gmeasure: 0.0000e+00 - val_loss: 0.6109 - val_binary_accuracy: 0.7000 - val_sensitivity: 1.0000 - val_specificity: 0.0000e+00 - val_gmeasure: 0.0000e+00
Epoch 2/10
640/640 [==============================] - 0s 132us/step - loss: 0.6140 - binary_accuracy: 0.6969 - sensitivity: 1.0000 - specificity: 0.0000e+00 - gmeasure: 0.0000e+00 - val_loss: 0.6112 - val_binary_accuracy: 0.7000 - val_sensitivity: 1.0000 - val_specificity: 0.0000e+00 - val_gmeasure: 0.0000e+00
Epoch 3/10
640/640 [==============================] - 0s 156us/step - loss: 0.6138 - binary_accuracy: 0.6969 - sensitivity: 1.0000 - specificity: 0.0000e+00 - gmeasure: 0.0000e+00 - val_loss: 0.6109 - val_binary_accuracy: 0.7000 - val_sensitivity: 1.0000 - val_specificity: 0.0000e+00 - val_gmeasure: 0.0000e+00
Epoch 4/10
640/640 [==============================] - 0s 127us/step - loss: 0.6136 - binary_accuracy: 0.6969 - sensitivity: 1.0000 - specificity: 0.0000e+00 - gmeasure: 0.0000e+00 - val_loss: 0.6109 - val_binary_accuracy: 0.7000 - val_sensitivity: 1.0000 - val_specificity: 0.0000e+00 - val_gmeasure: 0.0000e+00
Epoch 5/10
640/640 [==============================] - 0s 167us/step - loss: 0.6142 - binary_accuracy: 0.6969 - sensitivity: 1.0000 - specificity: 0.0000e+00 - gmeasure: 0.0000e+00 - val_loss: 0.6109 - val_binary_accuracy: 0.7000 - val_sensitivity: 1.0000 - val_specificity: 0.0000e+00 - val_gmeasure: 0.0000e+00
Epoch 6/10
640/640 [==============================] - 0s 146us/step - loss: 0.6135 - binary_accuracy: 0.6969 - sensitivity: 1.0000 - specificity: 0.0000e+00 - gmeasure: 0.0000e+00 - val_loss: 0.6109 - val_binary_accuracy: 0.7000 - val_sensitivity: 1.0000 - val_specificity: 0.0000e+00 - val_gmeasure: 0.0000e+00
Epoch 7/10
640/640 [==============================] - 0s 136us/step - loss: 0.6139 - binary_accuracy: 0.6969 - sensitivity: 1.0000 - specificity: 0.0000e+00 - gmeasure: 0.0000e+00 - val_loss: 0.6115 - val_binary_accuracy: 0.7000 - val_sensitivity: 1.0000 - val_specificity: 0.0000e+00 - val_gmeasure: 0.0000e+00
Epoch 8/10
640/640 [==============================] - 0s 100us/step - loss: 0.6146 - binary_accuracy: 0.6969 - sensitivity: 1.0000 - specificity: 0.0000e+00 - gmeasure: 0.0000e+00 - val_loss: 0.6110 - val_binary_accuracy: 0.7000 - val_sensitivity: 1.0000 - val_specificity: 0.0000e+00 - val_gmeasure: 0.0000e+00
Epoch 9/10
640/640 [==============================] - 0s 124us/step - loss: 0.6138 - binary_accuracy: 0.6969 - sensitivity: 1.0000 - specificity: 0.0000e+00 - gmeasure: 0.0000e+00 - val_loss: 0.6109 - val_binary_accuracy: 0.7000 - val_sensitivity: 1.0000 - val_specificity: 0.0000e+00 - val_gmeasure: 0.0000e+00
Epoch 10/10
640/640 [==============================] - 0s 118us/step - loss: 0.6135 - binary_accuracy: 0.6969 - sensitivity: 1.0000 - specificity: 0.0000e+00 - gmeasure: 0.0000e+00 - val_loss: 0.6110 - val_binary_accuracy: 0.7000 - val_sensitivity: 1.0000 - val_specificity: 0.0000e+00 - val_gmeasure: 0.0000e+00
[root |INFO|build_network.py:87] Load trained model weight at ./example_result/weight_2.h5
[root |INFO|build_network.py:147] Training end with time 2.8092546463012695!
[root |INFO|build_network.py:83] Saved trained model weight at ./example_result/weight_2.h5
[root |DEBUG|deepbiome.py:185] Save weight at ./example_result/weight_2.h5
[root |DEBUG|deepbiome.py:188] Save history at ./example_result/hist_2.json
[root |INFO|build_network.py:173] Evaluation start!
800/800 [==============================] - 0s 3us/step
[root |INFO|build_network.py:178] Evaluation end with time 0.008527040481567383!
[root |INFO|build_network.py:179] Evaluation: [0.6130011081695557, 0.6974999904632568, 1.0, 0.0, 0.0]
[root |INFO|build_network.py:173] Evaluation start!
200/200 [==============================] - 0s 12us/step
[root |INFO|build_network.py:178] Evaluation end with time 0.008930444717407227!
[root |INFO|build_network.py:179] Evaluation: [0.6493600606918335, 0.6549999713897705, 1.0, 0.0, 0.0]
[root |INFO|deepbiome.py:199] Compute time : 3.428178071975708
[root |INFO|deepbiome.py:200] 3 fold computing end!---------------------------------------------
[root |INFO|deepbiome.py:153] -------4 simulation start!----------------------------------
[root |INFO|readers.py:58] -----------------------------------------------------------------------
[root |INFO|readers.py:59] Construct Dataset
[root |INFO|readers.py:60] -----------------------------------------------------------------------
[root |INFO|readers.py:61] Load data
[root |INFO|deepbiome.py:164] -----------------------------------------------------------------
[root |INFO|deepbiome.py:165] Build network for 4 simulation
[root |INFO|build_network.py:521] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:522] Read phylogenetic tree information from /DATA/home/muha/github_repos/deepbiome/deepbiome/tests/data/genus48_dic.csv
[root |INFO|build_network.py:528] Phylogenetic tree level list: ['Genus', 'Family', 'Order', 'Class', 'Phylum']
[root |INFO|build_network.py:529] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:537] Genus: 48
[root |INFO|build_network.py:537] Family: 40
[root |INFO|build_network.py:537] Order: 23
[root |INFO|build_network.py:537] Class: 17
[root |INFO|build_network.py:537] Phylum: 9
[root |INFO|build_network.py:546] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:547] Phylogenetic_tree_dict info: ['Genus', 'Order', 'Class', 'Family', 'Phylum', 'Number']
[root |INFO|build_network.py:548] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:558] Build edge weights between [ Genus, Family]
[root |INFO|build_network.py:558] Build edge weights between [Family, Order]
[root |INFO|build_network.py:558] Build edge weights between [ Order, Class]
[root |INFO|build_network.py:558] Build edge weights between [ Class, Phylum]
[root |INFO|build_network.py:571] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:586] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:587] Build network based on phylogenetic tree information
[root |INFO|build_network.py:588] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:670] ------------------------------------------------------------------------------------------
Model: "model_1"
_________________________________________________________________
Layer (type) Output Shape Param #
=================================================================
input (InputLayer) (None, 48) 0
_________________________________________________________________
l1_dense (Dense_with_tree) (None, 40) 1960
_________________________________________________________________
l1_activation (Activation) (None, 40) 0
_________________________________________________________________
l2_dense (Dense_with_tree) (None, 23) 943
_________________________________________________________________
l2_activation (Activation) (None, 23) 0
_________________________________________________________________
l3_dense (Dense_with_tree) (None, 17) 408
_________________________________________________________________
l3_activation (Activation) (None, 17) 0
_________________________________________________________________
l4_dense (Dense_with_tree) (None, 9) 162
_________________________________________________________________
l4_activation (Activation) (None, 9) 0
_________________________________________________________________
last_dense_h (Dense) (None, 1) 10
_________________________________________________________________
p_hat (Activation) (None, 1) 0
=================================================================
Total params: 3,483
Trainable params: 3,483
Non-trainable params: 0
_________________________________________________________________
[root |INFO|build_network.py:61] Build Network
[root |INFO|build_network.py:62] Optimizer = adam
[root |INFO|build_network.py:63] Loss = binary_crossentropy
[root |INFO|build_network.py:64] Metrics = binary_accuracy, sensitivity, specificity, gmeasure
[root |INFO|build_network.py:87] Load trained model weight at ./example_result/weight_3.h5
[root |INFO|deepbiome.py:176] -----------------------------------------------------------------
[root |INFO|deepbiome.py:177] 4 fold computing start!----------------------------------
[root |INFO|build_network.py:137] Training start!
Train on 640 samples, validate on 160 samples
Epoch 1/10
640/640 [==============================] - 1s 1ms/step - loss: 0.5932 - binary_accuracy: 0.6859 - sensitivity: 1.0000 - specificity: 0.0000e+00 - gmeasure: 0.0000e+00 - val_loss: 0.5896 - val_binary_accuracy: 0.6875 - val_sensitivity: 1.0000 - val_specificity: 0.0000e+00 - val_gmeasure: 0.0000e+00
Epoch 2/10
640/640 [==============================] - 0s 502us/step - loss: 0.5770 - binary_accuracy: 0.6859 - sensitivity: 1.0000 - specificity: 0.0000e+00 - gmeasure: 0.0000e+00 - val_loss: 0.5793 - val_binary_accuracy: 0.6875 - val_sensitivity: 1.0000 - val_specificity: 0.0000e+00 - val_gmeasure: 0.0000e+00
Epoch 3/10
640/640 [==============================] - 0s 146us/step - loss: 0.5591 - binary_accuracy: 0.6859 - sensitivity: 1.0000 - specificity: 0.0000e+00 - gmeasure: 0.0000e+00 - val_loss: 0.5682 - val_binary_accuracy: 0.6875 - val_sensitivity: 1.0000 - val_specificity: 0.0000e+00 - val_gmeasure: 0.0000e+00
Epoch 4/10
640/640 [==============================] - 0s 422us/step - loss: 0.5451 - binary_accuracy: 0.6859 - sensitivity: 1.0000 - specificity: 0.0000e+00 - gmeasure: 0.0000e+00 - val_loss: 0.5566 - val_binary_accuracy: 0.6875 - val_sensitivity: 1.0000 - val_specificity: 0.0000e+00 - val_gmeasure: 0.0000e+00
Epoch 5/10
640/640 [==============================] - 0s 402us/step - loss: 0.5403 - binary_accuracy: 0.6859 - sensitivity: 1.0000 - specificity: 0.0000e+00 - gmeasure: 0.0000e+00 - val_loss: 0.5552 - val_binary_accuracy: 0.6875 - val_sensitivity: 1.0000 - val_specificity: 0.0000e+00 - val_gmeasure: 0.0000e+00
Epoch 6/10
640/640 [==============================] - 0s 158us/step - loss: 0.5326 - binary_accuracy: 0.6859 - sensitivity: 1.0000 - specificity: 0.0000e+00 - gmeasure: 0.0000e+00 - val_loss: 0.5409 - val_binary_accuracy: 0.6875 - val_sensitivity: 1.0000 - val_specificity: 0.0000e+00 - val_gmeasure: 0.0000e+00
Epoch 7/10
640/640 [==============================] - 0s 399us/step - loss: 0.5136 - binary_accuracy: 0.7203 - sensitivity: 1.0000 - specificity: 0.1122 - gmeasure: 0.2577 - val_loss: 0.5357 - val_binary_accuracy: 0.6875 - val_sensitivity: 1.0000 - val_specificity: 0.0000e+00 - val_gmeasure: 0.0000e+00
Epoch 8/10
640/640 [==============================] - 0s 315us/step - loss: 0.5015 - binary_accuracy: 0.7063 - sensitivity: 1.0000 - specificity: 0.0668 - gmeasure: 0.1809 - val_loss: 0.5221 - val_binary_accuracy: 0.7250 - val_sensitivity: 1.0000 - val_specificity: 0.0944 - val_gmeasure: 0.2541
Epoch 9/10
640/640 [==============================] - 0s 190us/step - loss: 0.4920 - binary_accuracy: 0.7219 - sensitivity: 1.0000 - specificity: 0.1111 - gmeasure: 0.2697 - val_loss: 0.5115 - val_binary_accuracy: 0.7125 - val_sensitivity: 1.0000 - val_specificity: 0.0588 - val_gmeasure: 0.1657
Epoch 10/10
640/640 [==============================] - 0s 448us/step - loss: 0.4800 - binary_accuracy: 0.7766 - sensitivity: 0.9957 - specificity: 0.3070 - gmeasure: 0.5399 - val_loss: 0.5035 - val_binary_accuracy: 0.7125 - val_sensitivity: 1.0000 - val_specificity: 0.0588 - val_gmeasure: 0.1657
[root |INFO|build_network.py:87] Load trained model weight at ./example_result/weight_3.h5
[root |INFO|build_network.py:147] Training end with time 4.435325384140015!
[root |INFO|build_network.py:83] Saved trained model weight at ./example_result/weight_3.h5
[root |DEBUG|deepbiome.py:185] Save weight at ./example_result/weight_3.h5
[root |DEBUG|deepbiome.py:188] Save history at ./example_result/hist_3.json
[root |INFO|build_network.py:173] Evaluation start!
800/800 [==============================] - 0s 4us/step
[root |INFO|build_network.py:178] Evaluation end with time 0.010611295700073242!
[root |INFO|build_network.py:179] Evaluation: [0.4794653058052063, 0.7200000286102295, 1.0, 0.10756972432136536, 0.3279782235622406]
[root |INFO|build_network.py:173] Evaluation start!
200/200 [==============================] - 0s 10us/step
[root |INFO|build_network.py:178] Evaluation end with time 0.008459091186523438!
[root |INFO|build_network.py:179] Evaluation: [0.45449164509773254, 0.7200000286102295, 1.0, 0.09677419066429138, 0.3110854923725128]
[root |INFO|deepbiome.py:199] Compute time : 5.07786226272583
[root |INFO|deepbiome.py:200] 4 fold computing end!---------------------------------------------
[root |INFO|deepbiome.py:153] -------5 simulation start!----------------------------------
[root |INFO|readers.py:58] -----------------------------------------------------------------------
[root |INFO|readers.py:59] Construct Dataset
[root |INFO|readers.py:60] -----------------------------------------------------------------------
[root |INFO|readers.py:61] Load data
[root |INFO|deepbiome.py:164] -----------------------------------------------------------------
[root |INFO|deepbiome.py:165] Build network for 5 simulation
[root |INFO|build_network.py:521] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:522] Read phylogenetic tree information from /DATA/home/muha/github_repos/deepbiome/deepbiome/tests/data/genus48_dic.csv
[root |INFO|build_network.py:528] Phylogenetic tree level list: ['Genus', 'Family', 'Order', 'Class', 'Phylum']
[root |INFO|build_network.py:529] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:537] Genus: 48
[root |INFO|build_network.py:537] Family: 40
[root |INFO|build_network.py:537] Order: 23
[root |INFO|build_network.py:537] Class: 17
[root |INFO|build_network.py:537] Phylum: 9
[root |INFO|build_network.py:546] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:547] Phylogenetic_tree_dict info: ['Genus', 'Order', 'Class', 'Family', 'Phylum', 'Number']
[root |INFO|build_network.py:548] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:558] Build edge weights between [ Genus, Family]
[root |INFO|build_network.py:558] Build edge weights between [Family, Order]
[root |INFO|build_network.py:558] Build edge weights between [ Order, Class]
[root |INFO|build_network.py:558] Build edge weights between [ Class, Phylum]
[root |INFO|build_network.py:571] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:586] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:587] Build network based on phylogenetic tree information
[root |INFO|build_network.py:588] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:670] ------------------------------------------------------------------------------------------
Model: "model_1"
_________________________________________________________________
Layer (type) Output Shape Param #
=================================================================
input (InputLayer) (None, 48) 0
_________________________________________________________________
l1_dense (Dense_with_tree) (None, 40) 1960
_________________________________________________________________
l1_activation (Activation) (None, 40) 0
_________________________________________________________________
l2_dense (Dense_with_tree) (None, 23) 943
_________________________________________________________________
l2_activation (Activation) (None, 23) 0
_________________________________________________________________
l3_dense (Dense_with_tree) (None, 17) 408
_________________________________________________________________
l3_activation (Activation) (None, 17) 0
_________________________________________________________________
l4_dense (Dense_with_tree) (None, 9) 162
_________________________________________________________________
l4_activation (Activation) (None, 9) 0
_________________________________________________________________
last_dense_h (Dense) (None, 1) 10
_________________________________________________________________
p_hat (Activation) (None, 1) 0
=================================================================
Total params: 3,483
Trainable params: 3,483
Non-trainable params: 0
_________________________________________________________________
[root |INFO|build_network.py:61] Build Network
[root |INFO|build_network.py:62] Optimizer = adam
[root |INFO|build_network.py:63] Loss = binary_crossentropy
[root |INFO|build_network.py:64] Metrics = binary_accuracy, sensitivity, specificity, gmeasure
[root |INFO|build_network.py:87] Load trained model weight at ./example_result/weight_4.h5
[root |INFO|deepbiome.py:176] -----------------------------------------------------------------
[root |INFO|deepbiome.py:177] 5 fold computing start!----------------------------------
[root |INFO|build_network.py:137] Training start!
Train on 640 samples, validate on 160 samples
Epoch 1/10
640/640 [==============================] - 0s 776us/step - loss: 0.6277 - binary_accuracy: 0.6797 - sensitivity: 1.0000 - specificity: 0.0000e+00 - gmeasure: 0.0000e+00 - val_loss: 0.6167 - val_binary_accuracy: 0.6938 - val_sensitivity: 1.0000 - val_specificity: 0.0000e+00 - val_gmeasure: 0.0000e+00
Epoch 2/10
640/640 [==============================] - 0s 123us/step - loss: 0.6281 - binary_accuracy: 0.6797 - sensitivity: 1.0000 - specificity: 0.0000e+00 - gmeasure: 0.0000e+00 - val_loss: 0.6162 - val_binary_accuracy: 0.6938 - val_sensitivity: 1.0000 - val_specificity: 0.0000e+00 - val_gmeasure: 0.0000e+00
Epoch 3/10
640/640 [==============================] - 0s 249us/step - loss: 0.6274 - binary_accuracy: 0.6797 - sensitivity: 1.0000 - specificity: 0.0000e+00 - gmeasure: 0.0000e+00 - val_loss: 0.6164 - val_binary_accuracy: 0.6938 - val_sensitivity: 1.0000 - val_specificity: 0.0000e+00 - val_gmeasure: 0.0000e+00
Epoch 4/10
640/640 [==============================] - 0s 444us/step - loss: 0.6275 - binary_accuracy: 0.6797 - sensitivity: 1.0000 - specificity: 0.0000e+00 - gmeasure: 0.0000e+00 - val_loss: 0.6163 - val_binary_accuracy: 0.6938 - val_sensitivity: 1.0000 - val_specificity: 0.0000e+00 - val_gmeasure: 0.0000e+00
Epoch 5/10
640/640 [==============================] - 0s 258us/step - loss: 0.6278 - binary_accuracy: 0.6797 - sensitivity: 1.0000 - specificity: 0.0000e+00 - gmeasure: 0.0000e+00 - val_loss: 0.6175 - val_binary_accuracy: 0.6938 - val_sensitivity: 1.0000 - val_specificity: 0.0000e+00 - val_gmeasure: 0.0000e+00
Epoch 6/10
640/640 [==============================] - 0s 130us/step - loss: 0.6274 - binary_accuracy: 0.6797 - sensitivity: 1.0000 - specificity: 0.0000e+00 - gmeasure: 0.0000e+00 - val_loss: 0.6163 - val_binary_accuracy: 0.6938 - val_sensitivity: 1.0000 - val_specificity: 0.0000e+00 - val_gmeasure: 0.0000e+00
Epoch 7/10
640/640 [==============================] - 0s 99us/step - loss: 0.6270 - binary_accuracy: 0.6797 - sensitivity: 1.0000 - specificity: 0.0000e+00 - gmeasure: 0.0000e+00 - val_loss: 0.6164 - val_binary_accuracy: 0.6938 - val_sensitivity: 1.0000 - val_specificity: 0.0000e+00 - val_gmeasure: 0.0000e+00
Epoch 8/10
640/640 [==============================] - 0s 96us/step - loss: 0.6271 - binary_accuracy: 0.6797 - sensitivity: 1.0000 - specificity: 0.0000e+00 - gmeasure: 0.0000e+00 - val_loss: 0.6163 - val_binary_accuracy: 0.6938 - val_sensitivity: 1.0000 - val_specificity: 0.0000e+00 - val_gmeasure: 0.0000e+00
Epoch 9/10
640/640 [==============================] - 0s 110us/step - loss: 0.6269 - binary_accuracy: 0.6797 - sensitivity: 1.0000 - specificity: 0.0000e+00 - gmeasure: 0.0000e+00 - val_loss: 0.6164 - val_binary_accuracy: 0.6938 - val_sensitivity: 1.0000 - val_specificity: 0.0000e+00 - val_gmeasure: 0.0000e+00
Epoch 10/10
640/640 [==============================] - 0s 121us/step - loss: 0.6267 - binary_accuracy: 0.6797 - sensitivity: 1.0000 - specificity: 0.0000e+00 - gmeasure: 0.0000e+00 - val_loss: 0.6166 - val_binary_accuracy: 0.6938 - val_sensitivity: 1.0000 - val_specificity: 0.0000e+00 - val_gmeasure: 0.0000e+00
[root |INFO|build_network.py:87] Load trained model weight at ./example_result/weight_4.h5
[root |INFO|build_network.py:147] Training end with time 2.8508307933807373!
[root |INFO|build_network.py:83] Saved trained model weight at ./example_result/weight_4.h5
[root |DEBUG|deepbiome.py:185] Save weight at ./example_result/weight_4.h5
[root |DEBUG|deepbiome.py:188] Save history at ./example_result/hist_4.json
[root |INFO|build_network.py:173] Evaluation start!
800/800 [==============================] - 0s 3us/step
[root |INFO|build_network.py:178] Evaluation end with time 0.008268117904663086!
[root |INFO|build_network.py:179] Evaluation: [0.6248783469200134, 0.6825000047683716, 1.0, 0.0, 0.0]
[root |INFO|build_network.py:173] Evaluation start!
200/200 [==============================] - 0s 9us/step
[root |INFO|build_network.py:178] Evaluation end with time 0.007416963577270508!
[root |INFO|build_network.py:179] Evaluation: [0.6074368953704834, 0.7049999833106995, 1.0, 0.0, 0.0]
[root |INFO|deepbiome.py:199] Compute time : 3.8774726390838623
[root |INFO|deepbiome.py:200] 5 fold computing end!---------------------------------------------
[root |INFO|deepbiome.py:211] -----------------------------------------------------------------
[root |INFO|deepbiome.py:212] Train Evaluation : ['loss' 'binary_accuracy' 'sensitivity' 'specificity' 'gmeasure']
[root |INFO|deepbiome.py:213] mean : [0.584 0.701 1.000 0.022 0.066]
[root |INFO|deepbiome.py:214] std : [0.053 0.016 0.000 0.043 0.131]
[root |INFO|deepbiome.py:215] -----------------------------------------------------------------
[root |INFO|deepbiome.py:216] Test Evaluation : ['loss' 'binary_accuracy' 'sensitivity' 'specificity' 'gmeasure']
[root |INFO|deepbiome.py:217] mean : [0.583 0.698 1.000 0.019 0.062]
[root |INFO|deepbiome.py:218] std : [0.067 0.024 0.000 0.039 0.124]
[root |INFO|deepbiome.py:219] -----------------------------------------------------------------
[root |INFO|deepbiome.py:230] Total Computing Ended
[root |INFO|deepbiome.py:231] -----------------------------------------------------------------
Let’s check the history plot again.
[25]:
with open('./%s/hist_0.json' % path_info['model_info']['model_dir'], 'r') as f:
history = json.load(f)
plt.plot(history['val_loss'], label='Validation')
plt.plot(history['loss'], label='Training')
plt.legend()
plt.xlabel('Epochs')
plt.ylabel('Loss')
plt.show()
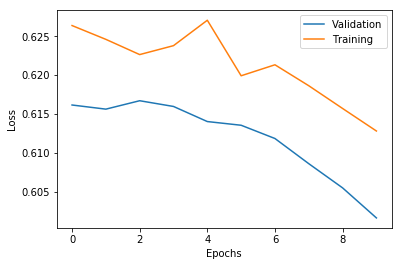
6. Load the pre-trained network for testing¶
To test the trained model, we can use the deepbiome_test function.
If you use the index file (idx_path), this function provides the evaluation using the test index (index set not included in the index file) for each fold. If not, this function provides the evaluation using the whole samples.
If number_of_fold is set to k, the function will test the model only with first k folds.
We can use the testing metrics different with the training. In the example below, we additionally used AUC metric.
[26]:
test_network_info = {
'architecture_info': {
'batch_normalization': 'False',
'drop_out': '0',
'weight_initial': 'glorot_uniform',
'weight_l1_penalty':'0.',
'weight_decay': 'phylogenetic_tree',
},
'model_info': {
'lr': '0.01',
'decay': '0.001',
'loss': 'binary_crossentropy',
'metrics': 'binary_accuracy, sensitivity, specificity, gmeasure, auc',
'taxa_selection_metrics': 'sensitivity, specificity, gmeasure, accuracy',
'network_class': 'DeepBiomeNetwork',
'optimizer': 'adam',
'reader_class': 'MicroBiomeClassificationReader',
'normalizer': 'normalize_minmax',
},
'test_info': {
'batch_size': 'None'
}
}
[27]:
test_path_info = {
'data_info': {
'data_path': resource_filename('deepbiome', 'tests/data'),
'idx_path': resource_filename('deepbiome', 'tests/data/onefile_idx.csv'),
'tree_info_path': resource_filename('deepbiome', 'tests/data/genus48_dic.csv'),
'x_path': 'onefile_x.csv',
'y_path': 'classification_y.csv'
},
'model_info': {
'evaluation': 'eval.npy',
'model_dir': './example_result/',
'weight': 'weight.h5'
}
}
[28]:
evaluation = deepbiome.deepbiome_test(log, test_network_info, test_path_info, number_of_fold=None)
[root |INFO|deepbiome.py:293] -----------------------------------------------------------------
[root |INFO|deepbiome.py:325] Test Evaluation : ['loss' 'binary_accuracy' 'sensitivity' 'specificity' 'gmeasure' 'auc']
[root |INFO|deepbiome.py:327] -------1 fold test start!----------------------------------
[root |INFO|readers.py:58] -----------------------------------------------------------------------
[root |INFO|readers.py:59] Construct Dataset
[root |INFO|readers.py:60] -----------------------------------------------------------------------
[root |INFO|readers.py:61] Load data
[root |INFO|deepbiome.py:338] -----------------------------------------------------------------
[root |INFO|deepbiome.py:339] Build network for 1 fold testing
[root |INFO|build_network.py:521] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:522] Read phylogenetic tree information from /DATA/home/muha/github_repos/deepbiome/deepbiome/tests/data/genus48_dic.csv
[root |INFO|build_network.py:528] Phylogenetic tree level list: ['Genus', 'Family', 'Order', 'Class', 'Phylum']
[root |INFO|build_network.py:529] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:537] Genus: 48
[root |INFO|build_network.py:537] Family: 40
[root |INFO|build_network.py:537] Order: 23
[root |INFO|build_network.py:537] Class: 17
[root |INFO|build_network.py:537] Phylum: 9
[root |INFO|build_network.py:546] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:547] Phylogenetic_tree_dict info: ['Genus', 'Order', 'Class', 'Family', 'Phylum', 'Number']
[root |INFO|build_network.py:548] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:558] Build edge weights between [ Genus, Family]
[root |INFO|build_network.py:558] Build edge weights between [Family, Order]
[root |INFO|build_network.py:558] Build edge weights between [ Order, Class]
[root |INFO|build_network.py:558] Build edge weights between [ Class, Phylum]
[root |INFO|build_network.py:571] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:586] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:587] Build network based on phylogenetic tree information
[root |INFO|build_network.py:588] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:670] ------------------------------------------------------------------------------------------
Model: "model_1"
_________________________________________________________________
Layer (type) Output Shape Param #
=================================================================
input (InputLayer) (None, 48) 0
_________________________________________________________________
l1_dense (Dense_with_tree) (None, 40) 1960
_________________________________________________________________
l1_activation (Activation) (None, 40) 0
_________________________________________________________________
l2_dense (Dense_with_tree) (None, 23) 943
_________________________________________________________________
l2_activation (Activation) (None, 23) 0
_________________________________________________________________
l3_dense (Dense_with_tree) (None, 17) 408
_________________________________________________________________
l3_activation (Activation) (None, 17) 0
_________________________________________________________________
l4_dense (Dense_with_tree) (None, 9) 162
_________________________________________________________________
l4_activation (Activation) (None, 9) 0
_________________________________________________________________
last_dense_h (Dense) (None, 1) 10
_________________________________________________________________
p_hat (Activation) (None, 1) 0
=================================================================
Total params: 3,483
Trainable params: 3,483
Non-trainable params: 0
_________________________________________________________________
[root |INFO|build_network.py:61] Build Network
[root |INFO|build_network.py:62] Optimizer = adam
[root |INFO|build_network.py:63] Loss = binary_crossentropy
[root |INFO|build_network.py:64] Metrics = binary_accuracy, sensitivity, specificity, gmeasure, auc
[root |INFO|build_network.py:87] Load trained model weight at ./example_result/weight_0.h5
[root |INFO|deepbiome.py:350] -----------------------------------------------------------------
[root |INFO|deepbiome.py:351] 1 fold computing start!----------------------------------
[root |INFO|build_network.py:173] Evaluation start!
200/200 [==============================] - 0s 594us/step
[root |INFO|build_network.py:178] Evaluation end with time 0.26632118225097656!
[root |INFO|build_network.py:179] Evaluation: [0.608977735042572, 0.6899999976158142, 1.0, 0.0, 0.0, 0.6397265195846558]
[root |INFO|deepbiome.py:356]
[root |INFO|deepbiome.py:357] Compute time : 0.8801772594451904
[root |INFO|deepbiome.py:358] 1 fold computing end!---------------------------------------------
[root |INFO|deepbiome.py:327] -------2 fold test start!----------------------------------
[root |INFO|readers.py:58] -----------------------------------------------------------------------
[root |INFO|readers.py:59] Construct Dataset
[root |INFO|readers.py:60] -----------------------------------------------------------------------
[root |INFO|readers.py:61] Load data
[root |INFO|deepbiome.py:338] -----------------------------------------------------------------
[root |INFO|deepbiome.py:339] Build network for 2 fold testing
[root |INFO|build_network.py:521] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:522] Read phylogenetic tree information from /DATA/home/muha/github_repos/deepbiome/deepbiome/tests/data/genus48_dic.csv
[root |INFO|build_network.py:528] Phylogenetic tree level list: ['Genus', 'Family', 'Order', 'Class', 'Phylum']
[root |INFO|build_network.py:529] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:537] Genus: 48
[root |INFO|build_network.py:537] Family: 40
[root |INFO|build_network.py:537] Order: 23
[root |INFO|build_network.py:537] Class: 17
[root |INFO|build_network.py:537] Phylum: 9
[root |INFO|build_network.py:546] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:547] Phylogenetic_tree_dict info: ['Genus', 'Order', 'Class', 'Family', 'Phylum', 'Number']
[root |INFO|build_network.py:548] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:558] Build edge weights between [ Genus, Family]
[root |INFO|build_network.py:558] Build edge weights between [Family, Order]
[root |INFO|build_network.py:558] Build edge weights between [ Order, Class]
[root |INFO|build_network.py:558] Build edge weights between [ Class, Phylum]
[root |INFO|build_network.py:571] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:586] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:587] Build network based on phylogenetic tree information
[root |INFO|build_network.py:588] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:670] ------------------------------------------------------------------------------------------
Model: "model_1"
_________________________________________________________________
Layer (type) Output Shape Param #
=================================================================
input (InputLayer) (None, 48) 0
_________________________________________________________________
l1_dense (Dense_with_tree) (None, 40) 1960
_________________________________________________________________
l1_activation (Activation) (None, 40) 0
_________________________________________________________________
l2_dense (Dense_with_tree) (None, 23) 943
_________________________________________________________________
l2_activation (Activation) (None, 23) 0
_________________________________________________________________
l3_dense (Dense_with_tree) (None, 17) 408
_________________________________________________________________
l3_activation (Activation) (None, 17) 0
_________________________________________________________________
l4_dense (Dense_with_tree) (None, 9) 162
_________________________________________________________________
l4_activation (Activation) (None, 9) 0
_________________________________________________________________
last_dense_h (Dense) (None, 1) 10
_________________________________________________________________
p_hat (Activation) (None, 1) 0
=================================================================
Total params: 3,483
Trainable params: 3,483
Non-trainable params: 0
_________________________________________________________________
[root |INFO|build_network.py:61] Build Network
[root |INFO|build_network.py:62] Optimizer = adam
[root |INFO|build_network.py:63] Loss = binary_crossentropy
[root |INFO|build_network.py:64] Metrics = binary_accuracy, sensitivity, specificity, gmeasure, auc
[root |INFO|build_network.py:87] Load trained model weight at ./example_result/weight_1.h5
[root |INFO|deepbiome.py:350] -----------------------------------------------------------------
[root |INFO|deepbiome.py:351] 2 fold computing start!----------------------------------
[root |INFO|build_network.py:173] Evaluation start!
200/200 [==============================] - 0s 542us/step
[root |INFO|build_network.py:178] Evaluation end with time 0.3015158176422119!
[root |INFO|build_network.py:179] Evaluation: [0.5931748747825623, 0.7200000286102295, 1.0, 0.0, 0.0, 0.4986979067325592]
[root |INFO|deepbiome.py:356]
[root |INFO|deepbiome.py:357] Compute time : 0.8466978073120117
[root |INFO|deepbiome.py:358] 2 fold computing end!---------------------------------------------
[root |INFO|deepbiome.py:327] -------3 fold test start!----------------------------------
[root |INFO|readers.py:58] -----------------------------------------------------------------------
[root |INFO|readers.py:59] Construct Dataset
[root |INFO|readers.py:60] -----------------------------------------------------------------------
[root |INFO|readers.py:61] Load data
[root |INFO|deepbiome.py:338] -----------------------------------------------------------------
[root |INFO|deepbiome.py:339] Build network for 3 fold testing
[root |INFO|build_network.py:521] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:522] Read phylogenetic tree information from /DATA/home/muha/github_repos/deepbiome/deepbiome/tests/data/genus48_dic.csv
[root |INFO|build_network.py:528] Phylogenetic tree level list: ['Genus', 'Family', 'Order', 'Class', 'Phylum']
[root |INFO|build_network.py:529] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:537] Genus: 48
[root |INFO|build_network.py:537] Family: 40
[root |INFO|build_network.py:537] Order: 23
[root |INFO|build_network.py:537] Class: 17
[root |INFO|build_network.py:537] Phylum: 9
[root |INFO|build_network.py:546] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:547] Phylogenetic_tree_dict info: ['Genus', 'Order', 'Class', 'Family', 'Phylum', 'Number']
[root |INFO|build_network.py:548] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:558] Build edge weights between [ Genus, Family]
[root |INFO|build_network.py:558] Build edge weights between [Family, Order]
[root |INFO|build_network.py:558] Build edge weights between [ Order, Class]
[root |INFO|build_network.py:558] Build edge weights between [ Class, Phylum]
[root |INFO|build_network.py:571] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:586] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:587] Build network based on phylogenetic tree information
[root |INFO|build_network.py:588] ------------------------------------------------------------------------------------------
Model: "model_1"
_________________________________________________________________
Layer (type) Output Shape Param #
=================================================================
[root |INFO|build_network.py:670] ------------------------------------------------------------------------------------------
input (InputLayer) (None, 48) 0
_________________________________________________________________
l1_dense (Dense_with_tree) (None, 40) 1960
_________________________________________________________________
l1_activation (Activation) (None, 40) 0
_________________________________________________________________
l2_dense (Dense_with_tree) (None, 23) 943
_________________________________________________________________
l2_activation (Activation) (None, 23) 0
_________________________________________________________________
l3_dense (Dense_with_tree) (None, 17) 408
_________________________________________________________________
l3_activation (Activation) (None, 17) 0
_________________________________________________________________
l4_dense (Dense_with_tree) (None, 9) 162
_________________________________________________________________
l4_activation (Activation) (None, 9) 0
_________________________________________________________________
last_dense_h (Dense) (None, 1) 10
_________________________________________________________________
p_hat (Activation) (None, 1) 0
=================================================================
Total params: 3,483
Trainable params: 3,483
Non-trainable params: 0
_________________________________________________________________
[root |INFO|build_network.py:61] Build Network
[root |INFO|build_network.py:62] Optimizer = adam
[root |INFO|build_network.py:63] Loss = binary_crossentropy
[root |INFO|build_network.py:64] Metrics = binary_accuracy, sensitivity, specificity, gmeasure, auc
[root |INFO|build_network.py:87] Load trained model weight at ./example_result/weight_2.h5
[root |INFO|deepbiome.py:350] -----------------------------------------------------------------
[root |INFO|deepbiome.py:351] 3 fold computing start!----------------------------------
[root |INFO|build_network.py:173] Evaluation start!
200/200 [==============================] - 0s 501us/step
[root |INFO|build_network.py:178] Evaluation end with time 0.2538328170776367!
[root |INFO|build_network.py:179] Evaluation: [0.6493600606918335, 0.6549999713897705, 1.0, 0.0, 0.0, 0.4984511435031891]
[root |INFO|deepbiome.py:356]
[root |INFO|deepbiome.py:357] Compute time : 0.9096362590789795
[root |INFO|deepbiome.py:358] 3 fold computing end!---------------------------------------------
[root |INFO|deepbiome.py:327] -------4 fold test start!----------------------------------
[root |INFO|readers.py:58] -----------------------------------------------------------------------
[root |INFO|readers.py:59] Construct Dataset
[root |INFO|readers.py:60] -----------------------------------------------------------------------
[root |INFO|readers.py:61] Load data
[root |INFO|deepbiome.py:338] -----------------------------------------------------------------
[root |INFO|deepbiome.py:339] Build network for 4 fold testing
[root |INFO|build_network.py:521] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:522] Read phylogenetic tree information from /DATA/home/muha/github_repos/deepbiome/deepbiome/tests/data/genus48_dic.csv
[root |INFO|build_network.py:528] Phylogenetic tree level list: ['Genus', 'Family', 'Order', 'Class', 'Phylum']
[root |INFO|build_network.py:529] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:537] Genus: 48
[root |INFO|build_network.py:537] Family: 40
[root |INFO|build_network.py:537] Order: 23
[root |INFO|build_network.py:537] Class: 17
[root |INFO|build_network.py:537] Phylum: 9
[root |INFO|build_network.py:546] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:547] Phylogenetic_tree_dict info: ['Genus', 'Order', 'Class', 'Family', 'Phylum', 'Number']
[root |INFO|build_network.py:548] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:558] Build edge weights between [ Genus, Family]
[root |INFO|build_network.py:558] Build edge weights between [Family, Order]
[root |INFO|build_network.py:558] Build edge weights between [ Order, Class]
[root |INFO|build_network.py:558] Build edge weights between [ Class, Phylum]
[root |INFO|build_network.py:571] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:586] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:587] Build network based on phylogenetic tree information
[root |INFO|build_network.py:588] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:670] ------------------------------------------------------------------------------------------
Model: "model_1"
_________________________________________________________________
Layer (type) Output Shape Param #
=================================================================
input (InputLayer) (None, 48) 0
_________________________________________________________________
l1_dense (Dense_with_tree) (None, 40) 1960
_________________________________________________________________
l1_activation (Activation) (None, 40) 0
_________________________________________________________________
l2_dense (Dense_with_tree) (None, 23) 943
_________________________________________________________________
l2_activation (Activation) (None, 23) 0
_________________________________________________________________
l3_dense (Dense_with_tree) (None, 17) 408
_________________________________________________________________
l3_activation (Activation) (None, 17) 0
_________________________________________________________________
l4_dense (Dense_with_tree) (None, 9) 162
_________________________________________________________________
l4_activation (Activation) (None, 9) 0
_________________________________________________________________
last_dense_h (Dense) (None, 1) 10
_________________________________________________________________
p_hat (Activation) (None, 1) 0
=================================================================
Total params: 3,483
Trainable params: 3,483
Non-trainable params: 0
_________________________________________________________________
[root |INFO|build_network.py:61] Build Network
[root |INFO|build_network.py:62] Optimizer = adam
[root |INFO|build_network.py:63] Loss = binary_crossentropy
[root |INFO|build_network.py:64] Metrics = binary_accuracy, sensitivity, specificity, gmeasure, auc
[root |INFO|build_network.py:87] Load trained model weight at ./example_result/weight_3.h5
[root |INFO|deepbiome.py:350] -----------------------------------------------------------------
[root |INFO|deepbiome.py:351] 4 fold computing start!----------------------------------
[root |INFO|build_network.py:173] Evaluation start!
200/200 [==============================] - 0s 432us/step
[root |INFO|build_network.py:178] Evaluation end with time 0.25353074073791504!
[root |INFO|build_network.py:179] Evaluation: [0.45449164509773254, 0.7200000286102295, 1.0, 0.09677419066429138, 0.3110854923725128, 0.8937587738037109]
[root |INFO|deepbiome.py:356]
[root |INFO|deepbiome.py:357] Compute time : 1.1399180889129639
[root |INFO|deepbiome.py:358] 4 fold computing end!---------------------------------------------
[root |INFO|deepbiome.py:327] -------5 fold test start!----------------------------------
[root |INFO|readers.py:58] -----------------------------------------------------------------------
[root |INFO|readers.py:59] Construct Dataset
[root |INFO|readers.py:60] -----------------------------------------------------------------------
[root |INFO|readers.py:61] Load data
[root |INFO|deepbiome.py:338] -----------------------------------------------------------------
[root |INFO|deepbiome.py:339] Build network for 5 fold testing
[root |INFO|build_network.py:521] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:522] Read phylogenetic tree information from /DATA/home/muha/github_repos/deepbiome/deepbiome/tests/data/genus48_dic.csv
[root |INFO|build_network.py:528] Phylogenetic tree level list: ['Genus', 'Family', 'Order', 'Class', 'Phylum']
[root |INFO|build_network.py:529] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:537] Genus: 48
[root |INFO|build_network.py:537] Family: 40
[root |INFO|build_network.py:537] Order: 23
[root |INFO|build_network.py:537] Class: 17
[root |INFO|build_network.py:537] Phylum: 9
[root |INFO|build_network.py:546] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:547] Phylogenetic_tree_dict info: ['Genus', 'Order', 'Class', 'Family', 'Phylum', 'Number']
[root |INFO|build_network.py:548] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:558] Build edge weights between [ Genus, Family]
[root |INFO|build_network.py:558] Build edge weights between [Family, Order]
[root |INFO|build_network.py:558] Build edge weights between [ Order, Class]
[root |INFO|build_network.py:558] Build edge weights between [ Class, Phylum]
[root |INFO|build_network.py:571] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:586] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:587] Build network based on phylogenetic tree information
[root |INFO|build_network.py:588] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:670] ------------------------------------------------------------------------------------------
Model: "model_1"
_________________________________________________________________
Layer (type) Output Shape Param #
=================================================================
input (InputLayer) (None, 48) 0
_________________________________________________________________
l1_dense (Dense_with_tree) (None, 40) 1960
_________________________________________________________________
l1_activation (Activation) (None, 40) 0
_________________________________________________________________
l2_dense (Dense_with_tree) (None, 23) 943
_________________________________________________________________
l2_activation (Activation) (None, 23) 0
_________________________________________________________________
l3_dense (Dense_with_tree) (None, 17) 408
_________________________________________________________________
l3_activation (Activation) (None, 17) 0
_________________________________________________________________
l4_dense (Dense_with_tree) (None, 9) 162
_________________________________________________________________
l4_activation (Activation) (None, 9) 0
_________________________________________________________________
last_dense_h (Dense) (None, 1) 10
_________________________________________________________________
p_hat (Activation) (None, 1) 0
=================================================================
Total params: 3,483
Trainable params: 3,483
Non-trainable params: 0
_________________________________________________________________
[root |INFO|build_network.py:61] Build Network
[root |INFO|build_network.py:62] Optimizer = adam
[root |INFO|build_network.py:63] Loss = binary_crossentropy
[root |INFO|build_network.py:64] Metrics = binary_accuracy, sensitivity, specificity, gmeasure, auc
[root |INFO|build_network.py:87] Load trained model weight at ./example_result/weight_4.h5
[root |INFO|deepbiome.py:350] -----------------------------------------------------------------
[root |INFO|deepbiome.py:351] 5 fold computing start!----------------------------------
[root |INFO|build_network.py:173] Evaluation start!
200/200 [==============================] - 0s 375us/step
[root |INFO|build_network.py:178] Evaluation end with time 0.25330519676208496!
[root |INFO|build_network.py:179] Evaluation: [0.6074368953704834, 0.7049999833106995, 1.0, 0.0, 0.0, 0.6676283478736877]
[root |INFO|deepbiome.py:356]
[root |INFO|deepbiome.py:357] Compute time : 0.8034312725067139
[root |INFO|deepbiome.py:358] 5 fold computing end!---------------------------------------------
[root |INFO|deepbiome.py:367] -----------------------------------------------------------------
[root |INFO|deepbiome.py:368] Test Evaluation : ['loss' 'binary_accuracy' 'sensitivity' 'specificity' 'gmeasure' 'auc']
[root |INFO|deepbiome.py:369] mean : [0.583 0.698 1.000 0.019 0.062 0.640]
[root |INFO|deepbiome.py:370] std : [0.067 0.024 0.000 0.039 0.124 0.145]
[root |INFO|deepbiome.py:371] -----------------------------------------------------------------
[root |INFO|deepbiome.py:372] Total Computing Ended
[root |INFO|deepbiome.py:373] -----------------------------------------------------------------
This function provides the evaluation result as a numpy array with a shape of (number of folds, number of evaluation measures).
[29]:
print(' %s' % ''.join(['%16s'%'loss']+ ['%16s'%s.strip() for s in test_network_info['model_info']['metrics'].split(',')]))
print('Mean: %s' % ''.join(['%16.4f'%v for v in np.mean(evaluation, axis=0)]))
print('Std : %s' % ''.join(['%16.4f'%v for v in np.std(evaluation, axis=0)]))
loss binary_accuracy sensitivity specificity gmeasure auc
Mean: 0.5827 0.6980 1.0000 0.0194 0.0622 0.6397
Std : 0.0668 0.0242 0.0000 0.0387 0.1244 0.1450
7. Load the pre-trained network for prediction¶
If you want to predict using the pre-trained model, you can use the deepbiome_prediction function. If number_of_fold is setted as k, the function will predict only with first k folds sample’s outputs.
If change_weight_for_each_fold is set as False, the function will predict the output of every folds by same weight from the given path. If change_weight_for_each_fold is set as True, the function will predict the output of by each fold weight.
If ‘get_y=True’, the function will provide a list of tuples (prediction, true output) as a output with the shape of (n_samples, 2, n_classes). If ‘get_y=False’, the function will provide predictions only. The output will have the shape of (n_samples, n_classes).
7.1 Prediction with fixed weight¶
If we want to predict new data from one pre-trained model, we can use the option below. We fixed the weight weight_0.h5 for predicting the whole samples from onefile_x.csv (without using index file).
[30]:
prediction_network_info = {
'architecture_info': {
'batch_normalization': 'False',
'drop_out': '0',
'weight_initial': 'glorot_uniform',
'weight_l1_penalty':'0.',
'weight_decay': 'phylogenetic_tree',
},
'model_info': {
'decay': '0.001',
'loss': 'binary_crossentropy',
'lr': '0.01',
'metrics': 'binary_accuracy, sensitivity, specificity, gmeasure, auc',
'network_class': 'DeepBiomeNetwork',
'normalizer': 'normalize_minmax',
'optimizer': 'adam',
'reader_class': 'MicroBiomeClassificationReader',
'taxa_selection_metrics': 'sensitivity, specificity, gmeasure, accuracy',
},
'test_info': {
'batch_size': 'None'
}
}
[31]:
prediction_path_info = {
'data_info': {
'data_path': resource_filename('deepbiome', 'tests/data'),
'tree_info_path': resource_filename('deepbiome', 'tests/data/genus48_dic.csv'),
'x_path': 'onefile_x.csv',
'y_path': 'classification_y.csv'
},
'model_info': {
'model_dir': './example_result/',
'weight': 'weight_0.h5'
}
}
[32]:
prediction = deepbiome.deepbiome_prediction(log, prediction_network_info, prediction_path_info,
num_classes = 1, number_of_fold=None)
[root |INFO|deepbiome.py:450] -----------------------------------------------------------------
[root |INFO|deepbiome.py:480] -------1 th repeatition prediction start!----------------------------------
[root |INFO|readers.py:58] -----------------------------------------------------------------------
[root |INFO|readers.py:59] Construct Dataset
[root |INFO|readers.py:60] -----------------------------------------------------------------------
[root |INFO|readers.py:61] Load data
[root |INFO|deepbiome.py:498] -----------------------------------------------------------------
[root |INFO|build_network.py:521] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:522] Read phylogenetic tree information from /DATA/home/muha/github_repos/deepbiome/deepbiome/tests/data/genus48_dic.csv
[root |INFO|build_network.py:528] Phylogenetic tree level list: ['Genus', 'Family', 'Order', 'Class', 'Phylum']
[root |INFO|build_network.py:529] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:537] Genus: 48
[root |INFO|build_network.py:537] Family: 40
[root |INFO|build_network.py:537] Order: 23
[root |INFO|build_network.py:537] Class: 17
[root |INFO|build_network.py:537] Phylum: 9
[root |INFO|build_network.py:546] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:547] Phylogenetic_tree_dict info: ['Genus', 'Order', 'Class', 'Family', 'Phylum', 'Number']
[root |INFO|build_network.py:548] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:558] Build edge weights between [ Genus, Family]
[root |INFO|build_network.py:558] Build edge weights between [Family, Order]
[root |INFO|build_network.py:558] Build edge weights between [ Order, Class]
[root |INFO|build_network.py:558] Build edge weights between [ Class, Phylum]
[root |INFO|build_network.py:571] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:586] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:587] Build network based on phylogenetic tree information
[root |INFO|build_network.py:588] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:670] ------------------------------------------------------------------------------------------
Model: "model_1"
_________________________________________________________________
Layer (type) Output Shape Param #
=================================================================
input (InputLayer) (None, 48) 0
_________________________________________________________________
l1_dense (Dense_with_tree) (None, 40) 1960
_________________________________________________________________
l1_activation (Activation) (None, 40) 0
_________________________________________________________________
l2_dense (Dense_with_tree) (None, 23) 943
_________________________________________________________________
l2_activation (Activation) (None, 23) 0
_________________________________________________________________
l3_dense (Dense_with_tree) (None, 17) 408
_________________________________________________________________
l3_activation (Activation) (None, 17) 0
_________________________________________________________________
l4_dense (Dense_with_tree) (None, 9) 162
_________________________________________________________________
l4_activation (Activation) (None, 9) 0
_________________________________________________________________
last_dense_h (Dense) (None, 1) 10
_________________________________________________________________
p_hat (Activation) (None, 1) 0
=================================================================
Total params: 3,483
Trainable params: 3,483
Non-trainable params: 0
_________________________________________________________________
[root |INFO|build_network.py:61] Build Network
[root |INFO|build_network.py:62] Optimizer = adam
[root |INFO|build_network.py:63] Loss = binary_crossentropy
[root |INFO|build_network.py:64] Metrics = binary_accuracy, sensitivity, specificity, gmeasure, auc
[root |INFO|build_network.py:87] Load trained model weight at ./example_result/weight_0.h5
[root |INFO|deepbiome.py:509] -----------------------------------------------------------------
[root |INFO|build_network.py:193] Prediction start!
1000/1000 [==============================] - 0s 51us/step
[root |INFO|build_network.py:198] Prediction end with time 0.05417037010192871!
[root |INFO|deepbiome.py:513] Compute time : 0.5956833362579346
[root |INFO|deepbiome.py:514] 1 fold computing end!---------------------------------------------
[root |INFO|deepbiome.py:519] Total Computing Ended
[root |INFO|deepbiome.py:520] -----------------------------------------------------------------
[33]:
prediction.shape
[33]:
(1, 1000, 1)
[34]:
prediction[0,:10]
[34]:
array([[0.684],
[0.688],
[0.676],
[0.662],
[0.674],
[0.739],
[0.688],
[0.670],
[0.669],
[0.667]], dtype=float32)
7.2 Prediction with each fold weight¶
If we want to predict the cross-validation outputs, we can use the option belows.
The example below shows how to predict the 5-fold cross-validation outputs. (If we use the leave-one-out-cross-validation (LOOCV) for training, we can predict the LOOCV outputs by same way.) We set idx_path for using the index file onefile_idx.csv to predict only the test set for each fold.
[35]:
prediction_network_info = {
'architecture_info': {
'batch_normalization': 'False',
'drop_out': '0',
'weight_initial': 'glorot_uniform',
'weight_l1_penalty':'0.',
'weight_decay': 'phylogenetic_tree',
},
'model_info': {
'decay': '0.001',
'loss': 'binary_crossentropy',
'lr': '0.01',
'metrics': 'binary_accuracy, sensitivity, specificity, gmeasure, auc',
'network_class': 'DeepBiomeNetwork',
'normalizer': 'normalize_minmax',
'optimizer': 'adam',
'reader_class': 'MicroBiomeClassificationReader',
'taxa_selection_metrics': 'sensitivity, specificity, gmeasure, accuracy',
},
'test_info': {
'batch_size': 'None'
}
}
[36]:
prediction_path_info = {
'data_info': {
'data_path': resource_filename('deepbiome', 'tests/data'),
'idx_path': resource_filename('deepbiome', 'tests/data/onefile_idx.csv'),
'tree_info_path': resource_filename('deepbiome', 'tests/data/genus48_dic.csv'),
'x_path': 'onefile_x.csv',
'y_path': 'classification_y.csv'
},
'model_info': {
'model_dir': './example_result/',
'weight': 'weight.h5'
}
}
To predict the CV outputs from each fold, we set change_weight_for_each_fold = True. Also, we set get_y=True to get the paired output of each prediction too.
[37]:
prediction = deepbiome.deepbiome_prediction(log, prediction_network_info, prediction_path_info,
num_classes = 1, number_of_fold=None,
change_weight_for_each_fold = True,
get_y=True)
[root |INFO|deepbiome.py:450] -----------------------------------------------------------------
[root |INFO|deepbiome.py:480] -------1 th repeatition prediction start!----------------------------------
[root |INFO|readers.py:58] -----------------------------------------------------------------------
[root |INFO|readers.py:59] Construct Dataset
[root |INFO|readers.py:60] -----------------------------------------------------------------------
[root |INFO|readers.py:61] Load data
[root |INFO|deepbiome.py:498] -----------------------------------------------------------------
[root |INFO|build_network.py:521] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:522] Read phylogenetic tree information from /DATA/home/muha/github_repos/deepbiome/deepbiome/tests/data/genus48_dic.csv
[root |INFO|build_network.py:528] Phylogenetic tree level list: ['Genus', 'Family', 'Order', 'Class', 'Phylum']
[root |INFO|build_network.py:529] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:537] Genus: 48
[root |INFO|build_network.py:537] Family: 40
[root |INFO|build_network.py:537] Order: 23
[root |INFO|build_network.py:537] Class: 17
[root |INFO|build_network.py:537] Phylum: 9
[root |INFO|build_network.py:546] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:547] Phylogenetic_tree_dict info: ['Genus', 'Order', 'Class', 'Family', 'Phylum', 'Number']
[root |INFO|build_network.py:548] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:558] Build edge weights between [ Genus, Family]
[root |INFO|build_network.py:558] Build edge weights between [Family, Order]
[root |INFO|build_network.py:558] Build edge weights between [ Order, Class]
[root |INFO|build_network.py:558] Build edge weights between [ Class, Phylum]
[root |INFO|build_network.py:571] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:586] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:587] Build network based on phylogenetic tree information
[root |INFO|build_network.py:588] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:670] ------------------------------------------------------------------------------------------
Model: "model_1"
_________________________________________________________________
Layer (type) Output Shape Param #
=================================================================
input (InputLayer) (None, 48) 0
_________________________________________________________________
l1_dense (Dense_with_tree) (None, 40) 1960
_________________________________________________________________
l1_activation (Activation) (None, 40) 0
_________________________________________________________________
l2_dense (Dense_with_tree) (None, 23) 943
_________________________________________________________________
l2_activation (Activation) (None, 23) 0
_________________________________________________________________
l3_dense (Dense_with_tree) (None, 17) 408
_________________________________________________________________
l3_activation (Activation) (None, 17) 0
_________________________________________________________________
l4_dense (Dense_with_tree) (None, 9) 162
_________________________________________________________________
l4_activation (Activation) (None, 9) 0
_________________________________________________________________
last_dense_h (Dense) (None, 1) 10
_________________________________________________________________
p_hat (Activation) (None, 1) 0
=================================================================
Total params: 3,483
Trainable params: 3,483
Non-trainable params: 0
_________________________________________________________________
[root |INFO|build_network.py:61] Build Network
[root |INFO|build_network.py:62] Optimizer = adam
[root |INFO|build_network.py:63] Loss = binary_crossentropy
[root |INFO|build_network.py:64] Metrics = binary_accuracy, sensitivity, specificity, gmeasure, auc
[root |INFO|build_network.py:87] Load trained model weight at ./example_result/weight_0.h5
[root |INFO|deepbiome.py:509] -----------------------------------------------------------------
[root |INFO|build_network.py:193] Prediction start!
200/200 [==============================] - 0s 147us/step
[root |INFO|build_network.py:198] Prediction end with time 0.032140254974365234!
[root |INFO|deepbiome.py:513] Compute time : 0.6093075275421143
[root |INFO|deepbiome.py:514] 1 fold computing end!---------------------------------------------
[root |INFO|deepbiome.py:480] -------2 th repeatition prediction start!----------------------------------
[root |INFO|readers.py:58] -----------------------------------------------------------------------
[root |INFO|readers.py:59] Construct Dataset
[root |INFO|readers.py:60] -----------------------------------------------------------------------
[root |INFO|readers.py:61] Load data
[root |INFO|deepbiome.py:498] -----------------------------------------------------------------
[root |INFO|build_network.py:521] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:522] Read phylogenetic tree information from /DATA/home/muha/github_repos/deepbiome/deepbiome/tests/data/genus48_dic.csv
[root |INFO|build_network.py:528] Phylogenetic tree level list: ['Genus', 'Family', 'Order', 'Class', 'Phylum']
[root |INFO|build_network.py:529] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:537] Genus: 48
[root |INFO|build_network.py:537] Family: 40
[root |INFO|build_network.py:537] Order: 23
[root |INFO|build_network.py:537] Class: 17
[root |INFO|build_network.py:537] Phylum: 9
[root |INFO|build_network.py:546] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:547] Phylogenetic_tree_dict info: ['Genus', 'Order', 'Class', 'Family', 'Phylum', 'Number']
[root |INFO|build_network.py:548] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:558] Build edge weights between [ Genus, Family]
[root |INFO|build_network.py:558] Build edge weights between [Family, Order]
[root |INFO|build_network.py:558] Build edge weights between [ Order, Class]
[root |INFO|build_network.py:558] Build edge weights between [ Class, Phylum]
[root |INFO|build_network.py:571] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:586] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:587] Build network based on phylogenetic tree information
[root |INFO|build_network.py:588] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:670] ------------------------------------------------------------------------------------------
Model: "model_2"
_________________________________________________________________
Layer (type) Output Shape Param #
=================================================================
input (InputLayer) (None, 48) 0
_________________________________________________________________
l1_dense (Dense_with_tree) (None, 40) 1960
_________________________________________________________________
l1_activation (Activation) (None, 40) 0
_________________________________________________________________
l2_dense (Dense_with_tree) (None, 23) 943
_________________________________________________________________
l2_activation (Activation) (None, 23) 0
_________________________________________________________________
l3_dense (Dense_with_tree) (None, 17) 408
_________________________________________________________________
l3_activation (Activation) (None, 17) 0
_________________________________________________________________
l4_dense (Dense_with_tree) (None, 9) 162
_________________________________________________________________
l4_activation (Activation) (None, 9) 0
_________________________________________________________________
last_dense_h (Dense) (None, 1) 10
_________________________________________________________________
p_hat (Activation) (None, 1) 0
=================================================================
Total params: 3,483
Trainable params: 3,483
Non-trainable params: 0
_________________________________________________________________
[root |INFO|build_network.py:61] Build Network
[root |INFO|build_network.py:62] Optimizer = adam
[root |INFO|build_network.py:63] Loss = binary_crossentropy
[root |INFO|build_network.py:64] Metrics = binary_accuracy, sensitivity, specificity, gmeasure, auc
[root |INFO|build_network.py:87] Load trained model weight at ./example_result/weight_1.h5
[root |INFO|deepbiome.py:509] -----------------------------------------------------------------
[root |INFO|build_network.py:193] Prediction start!
200/200 [==============================] - 0s 219us/step
[root |INFO|build_network.py:198] Prediction end with time 0.0466768741607666!
[root |INFO|deepbiome.py:513] Compute time : 0.6837012767791748
[root |INFO|deepbiome.py:514] 2 fold computing end!---------------------------------------------
[root |INFO|deepbiome.py:480] -------3 th repeatition prediction start!----------------------------------
[root |INFO|readers.py:58] -----------------------------------------------------------------------
[root |INFO|readers.py:59] Construct Dataset
[root |INFO|readers.py:60] -----------------------------------------------------------------------
[root |INFO|readers.py:61] Load data
[root |INFO|deepbiome.py:498] -----------------------------------------------------------------
[root |INFO|build_network.py:521] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:522] Read phylogenetic tree information from /DATA/home/muha/github_repos/deepbiome/deepbiome/tests/data/genus48_dic.csv
[root |INFO|build_network.py:528] Phylogenetic tree level list: ['Genus', 'Family', 'Order', 'Class', 'Phylum']
[root |INFO|build_network.py:529] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:537] Genus: 48
[root |INFO|build_network.py:537] Family: 40
[root |INFO|build_network.py:537] Order: 23
[root |INFO|build_network.py:537] Class: 17
[root |INFO|build_network.py:537] Phylum: 9
[root |INFO|build_network.py:546] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:547] Phylogenetic_tree_dict info: ['Genus', 'Order', 'Class', 'Family', 'Phylum', 'Number']
[root |INFO|build_network.py:548] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:558] Build edge weights between [ Genus, Family]
[root |INFO|build_network.py:558] Build edge weights between [Family, Order]
[root |INFO|build_network.py:558] Build edge weights between [ Order, Class]
[root |INFO|build_network.py:558] Build edge weights between [ Class, Phylum]
[root |INFO|build_network.py:571] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:586] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:587] Build network based on phylogenetic tree information
[root |INFO|build_network.py:588] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:670] ------------------------------------------------------------------------------------------
Model: "model_3"
_________________________________________________________________
Layer (type) Output Shape Param #
=================================================================
input (InputLayer) (None, 48) 0
_________________________________________________________________
l1_dense (Dense_with_tree) (None, 40) 1960
_________________________________________________________________
l1_activation (Activation) (None, 40) 0
_________________________________________________________________
l2_dense (Dense_with_tree) (None, 23) 943
_________________________________________________________________
l2_activation (Activation) (None, 23) 0
_________________________________________________________________
l3_dense (Dense_with_tree) (None, 17) 408
_________________________________________________________________
l3_activation (Activation) (None, 17) 0
_________________________________________________________________
l4_dense (Dense_with_tree) (None, 9) 162
_________________________________________________________________
l4_activation (Activation) (None, 9) 0
_________________________________________________________________
last_dense_h (Dense) (None, 1) 10
_________________________________________________________________
p_hat (Activation) (None, 1) 0
=================================================================
Total params: 3,483
Trainable params: 3,483
Non-trainable params: 0
_________________________________________________________________
[root |INFO|build_network.py:61] Build Network
[root |INFO|build_network.py:62] Optimizer = adam
[root |INFO|build_network.py:63] Loss = binary_crossentropy
[root |INFO|build_network.py:64] Metrics = binary_accuracy, sensitivity, specificity, gmeasure, auc
[root |INFO|build_network.py:87] Load trained model weight at ./example_result/weight_2.h5
[root |INFO|deepbiome.py:509] -----------------------------------------------------------------
[root |INFO|build_network.py:193] Prediction start!
200/200 [==============================] - 0s 282us/step
[root |INFO|build_network.py:198] Prediction end with time 0.05935072898864746!
[root |INFO|deepbiome.py:513] Compute time : 0.673020601272583
[root |INFO|deepbiome.py:514] 3 fold computing end!---------------------------------------------
[root |INFO|deepbiome.py:480] -------4 th repeatition prediction start!----------------------------------
[root |INFO|readers.py:58] -----------------------------------------------------------------------
[root |INFO|readers.py:59] Construct Dataset
[root |INFO|readers.py:60] -----------------------------------------------------------------------
[root |INFO|readers.py:61] Load data
[root |INFO|deepbiome.py:498] -----------------------------------------------------------------
[root |INFO|build_network.py:521] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:522] Read phylogenetic tree information from /DATA/home/muha/github_repos/deepbiome/deepbiome/tests/data/genus48_dic.csv
[root |INFO|build_network.py:528] Phylogenetic tree level list: ['Genus', 'Family', 'Order', 'Class', 'Phylum']
[root |INFO|build_network.py:529] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:537] Genus: 48
[root |INFO|build_network.py:537] Family: 40
[root |INFO|build_network.py:537] Order: 23
[root |INFO|build_network.py:537] Class: 17
[root |INFO|build_network.py:537] Phylum: 9
[root |INFO|build_network.py:546] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:547] Phylogenetic_tree_dict info: ['Genus', 'Order', 'Class', 'Family', 'Phylum', 'Number']
[root |INFO|build_network.py:548] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:558] Build edge weights between [ Genus, Family]
[root |INFO|build_network.py:558] Build edge weights between [Family, Order]
[root |INFO|build_network.py:558] Build edge weights between [ Order, Class]
[root |INFO|build_network.py:558] Build edge weights between [ Class, Phylum]
[root |INFO|build_network.py:571] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:586] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:587] Build network based on phylogenetic tree information
[root |INFO|build_network.py:588] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:670] ------------------------------------------------------------------------------------------
Model: "model_4"
_________________________________________________________________
Layer (type) Output Shape Param #
=================================================================
input (InputLayer) (None, 48) 0
_________________________________________________________________
l1_dense (Dense_with_tree) (None, 40) 1960
_________________________________________________________________
l1_activation (Activation) (None, 40) 0
_________________________________________________________________
l2_dense (Dense_with_tree) (None, 23) 943
_________________________________________________________________
l2_activation (Activation) (None, 23) 0
_________________________________________________________________
l3_dense (Dense_with_tree) (None, 17) 408
_________________________________________________________________
l3_activation (Activation) (None, 17) 0
_________________________________________________________________
l4_dense (Dense_with_tree) (None, 9) 162
_________________________________________________________________
l4_activation (Activation) (None, 9) 0
_________________________________________________________________
last_dense_h (Dense) (None, 1) 10
_________________________________________________________________
p_hat (Activation) (None, 1) 0
=================================================================
Total params: 3,483
Trainable params: 3,483
Non-trainable params: 0
_________________________________________________________________
[root |INFO|build_network.py:61] Build Network
[root |INFO|build_network.py:62] Optimizer = adam
[root |INFO|build_network.py:63] Loss = binary_crossentropy
[root |INFO|build_network.py:64] Metrics = binary_accuracy, sensitivity, specificity, gmeasure, auc
[root |INFO|build_network.py:87] Load trained model weight at ./example_result/weight_3.h5
[root |INFO|deepbiome.py:509] -----------------------------------------------------------------
[root |INFO|build_network.py:193] Prediction start!
200/200 [==============================] - 0s 516us/step
[root |INFO|build_network.py:198] Prediction end with time 0.10687398910522461!
[root |INFO|deepbiome.py:513] Compute time : 0.9963078498840332
[root |INFO|deepbiome.py:514] 4 fold computing end!---------------------------------------------
[root |INFO|deepbiome.py:480] -------5 th repeatition prediction start!----------------------------------
[root |INFO|readers.py:58] -----------------------------------------------------------------------
[root |INFO|readers.py:59] Construct Dataset
[root |INFO|readers.py:60] -----------------------------------------------------------------------
[root |INFO|readers.py:61] Load data
[root |INFO|deepbiome.py:498] -----------------------------------------------------------------
[root |INFO|build_network.py:521] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:522] Read phylogenetic tree information from /DATA/home/muha/github_repos/deepbiome/deepbiome/tests/data/genus48_dic.csv
[root |INFO|build_network.py:528] Phylogenetic tree level list: ['Genus', 'Family', 'Order', 'Class', 'Phylum']
[root |INFO|build_network.py:529] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:537] Genus: 48
[root |INFO|build_network.py:537] Family: 40
[root |INFO|build_network.py:537] Order: 23
[root |INFO|build_network.py:537] Class: 17
[root |INFO|build_network.py:537] Phylum: 9
[root |INFO|build_network.py:546] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:547] Phylogenetic_tree_dict info: ['Genus', 'Order', 'Class', 'Family', 'Phylum', 'Number']
[root |INFO|build_network.py:548] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:558] Build edge weights between [ Genus, Family]
[root |INFO|build_network.py:558] Build edge weights between [Family, Order]
[root |INFO|build_network.py:558] Build edge weights between [ Order, Class]
[root |INFO|build_network.py:558] Build edge weights between [ Class, Phylum]
[root |INFO|build_network.py:571] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:586] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:587] Build network based on phylogenetic tree information
[root |INFO|build_network.py:588] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:670] ------------------------------------------------------------------------------------------
Model: "model_5"
_________________________________________________________________
Layer (type) Output Shape Param #
=================================================================
input (InputLayer) (None, 48) 0
_________________________________________________________________
l1_dense (Dense_with_tree) (None, 40) 1960
_________________________________________________________________
l1_activation (Activation) (None, 40) 0
_________________________________________________________________
l2_dense (Dense_with_tree) (None, 23) 943
_________________________________________________________________
l2_activation (Activation) (None, 23) 0
_________________________________________________________________
l3_dense (Dense_with_tree) (None, 17) 408
_________________________________________________________________
l3_activation (Activation) (None, 17) 0
_________________________________________________________________
l4_dense (Dense_with_tree) (None, 9) 162
_________________________________________________________________
l4_activation (Activation) (None, 9) 0
_________________________________________________________________
last_dense_h (Dense) (None, 1) 10
_________________________________________________________________
p_hat (Activation) (None, 1) 0
=================================================================
Total params: 3,483
Trainable params: 3,483
Non-trainable params: 0
_________________________________________________________________
[root |INFO|build_network.py:61] Build Network
[root |INFO|build_network.py:62] Optimizer = adam
[root |INFO|build_network.py:63] Loss = binary_crossentropy
[root |INFO|build_network.py:64] Metrics = binary_accuracy, sensitivity, specificity, gmeasure, auc
[root |INFO|build_network.py:87] Load trained model weight at ./example_result/weight_4.h5
[root |INFO|deepbiome.py:509] -----------------------------------------------------------------
[root |INFO|build_network.py:193] Prediction start!
200/200 [==============================] - 0s 723us/step
[root |INFO|build_network.py:198] Prediction end with time 0.1472012996673584!
[root |INFO|deepbiome.py:513] Compute time : 0.9382455348968506
[root |INFO|deepbiome.py:514] 5 fold computing end!---------------------------------------------
[root |INFO|deepbiome.py:519] Total Computing Ended
[root |INFO|deepbiome.py:520] -----------------------------------------------------------------
We gathered the outputs from each fold.
[38]:
prediction = np.vstack(prediction)
Since we set the option get_y=True, the output has the shape of (n_samples, 2, n_classes). With this options, we can get the CV predictions and the true output of each predictions.
Now, we can calculate the CV performance by the CV predictions.
[39]:
predict_output = prediction[:,0]
true_output = prediction[:,1]
log.info('Shape of the predict function output: %s' % str(prediction.shape))
log.info('Shape of the prediction: %s' % str(predict_output.shape))
log.info('Shape of the true_output for each prediction: %s' % str(true_output.shape))
[root |INFO|<ipython-input-39-edc88a58497e>:4] Shape of the predict function output: (1000, 2, 1)
[root |INFO|<ipython-input-39-edc88a58497e>:5] Shape of the prediction: (1000, 1)
[root |INFO|<ipython-input-39-edc88a58497e>:6] Shape of the true_output for each prediction: (1000, 1)
[40]:
log.info('CV accuracy: %6.3f' % np.mean((predict_output >= 0.5) == true_output))
[root |INFO|<ipython-input-40-ecaee2413087>:1] CV accuracy: 0.698
8. Load trained weight matrix¶
The deepbiome_get_trained_weight function convert the trained weight *.h5 saved from the deepbiome_train to a list of pandas dataframe. In this exampe, the list has numpy array of weights from 6 layers. ([genus to family, family to order, order to Class, class to phylum, phylum to output])
[41]:
weight_path = '%s/%s' % (path_info['model_info']['model_dir'], 'weight_0.h5')
trained_weight_list = deepbiome.deepbiome_get_trained_weight(log, network_info, path_info, num_classes=1, weight_path=weight_path)
log.info(len(trained_weight_list))
[root |INFO|readers.py:58] -----------------------------------------------------------------------
[root |INFO|readers.py:59] Construct Dataset
[root |INFO|readers.py:60] -----------------------------------------------------------------------
[root |INFO|readers.py:61] Load data
[root |INFO|build_network.py:521] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:522] Read phylogenetic tree information from /DATA/home/muha/github_repos/deepbiome/deepbiome/tests/data/genus48_dic.csv
[root |INFO|build_network.py:528] Phylogenetic tree level list: ['Genus', 'Family', 'Order', 'Class', 'Phylum']
[root |INFO|build_network.py:529] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:537] Genus: 48
[root |INFO|build_network.py:537] Family: 40
[root |INFO|build_network.py:537] Order: 23
[root |INFO|build_network.py:537] Class: 17
[root |INFO|build_network.py:537] Phylum: 9
[root |INFO|build_network.py:546] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:547] Phylogenetic_tree_dict info: ['Genus', 'Order', 'Class', 'Family', 'Phylum', 'Number']
[root |INFO|build_network.py:548] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:558] Build edge weights between [ Genus, Family]
[root |INFO|build_network.py:558] Build edge weights between [Family, Order]
[root |INFO|build_network.py:558] Build edge weights between [ Order, Class]
[root |INFO|build_network.py:558] Build edge weights between [ Class, Phylum]
[root |INFO|build_network.py:571] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:586] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:587] Build network based on phylogenetic tree information
[root |INFO|build_network.py:588] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:670] ------------------------------------------------------------------------------------------
Model: "model_1"
_________________________________________________________________
Layer (type) Output Shape Param #
=================================================================
input (InputLayer) (None, 48) 0
_________________________________________________________________
l1_dense (Dense_with_tree) (None, 40) 1960
_________________________________________________________________
l1_activation (Activation) (None, 40) 0
_________________________________________________________________
l2_dense (Dense_with_tree) (None, 23) 943
_________________________________________________________________
l2_activation (Activation) (None, 23) 0
_________________________________________________________________
l3_dense (Dense_with_tree) (None, 17) 408
_________________________________________________________________
l3_activation (Activation) (None, 17) 0
_________________________________________________________________
l4_dense (Dense_with_tree) (None, 9) 162
_________________________________________________________________
l4_activation (Activation) (None, 9) 0
_________________________________________________________________
last_dense_h (Dense) (None, 1) 10
_________________________________________________________________
p_hat (Activation) (None, 1) 0
=================================================================
Total params: 3,483
Trainable params: 3,483
Non-trainable params: 0
_________________________________________________________________
[root |INFO|<ipython-input-41-9505ee8dcaa8>:3] 5
First weight between the genus and family layers has the shape of (number of genus = 48, number of family = 40)
[42]:
log.info(trained_weight_list[0].shape)
[root |INFO|<ipython-input-42-c71fa46ab178>:1] (48, 40)
[43]:
trained_weight_list[0]
[43]:
| Streptococcaceae | Cellulomonadaceae | Veillonellaceae | Actinomycetaceae | Flavobacteriaceae | Prevotellaceae | Porphyromonadaceae | Clostridiales_Incertae_Sedis_XI | Fusobacteriaceae | Propionibacteriaceae | ... | Methylobacteriaceae | OD1_genera_incertae_sedis | Microbacteriaceae | Sphingomonadaceae | Bacteroidaceae | Bdellovibrionaceae | Chloroplast | Rhodobacteraceae | Ruminococcaceae | Pseudonocardiaceae | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Streptococcus | -0.174 | -0.001 | -0.003 | -0.002 | 0.001 | -0.001 | -0.000 | 0.002 | 0.002 | 0.002 | ... | 0.001 | -0.002 | 0.001 | 0.000 | -0.003 | 0.000 | 0.003 | 0.002 | -0.002 | 0.000 |
| Tropheryma | -0.003 | -0.248 | 0.001 | 0.001 | -0.002 | -0.001 | 0.002 | 0.003 | 0.006 | 0.005 | ... | -0.002 | -0.001 | 0.002 | -0.001 | -0.007 | -0.001 | 0.003 | 0.002 | -0.002 | -0.001 |
| Veillonella | -0.001 | -0.002 | -0.205 | 0.002 | -0.000 | 0.002 | -0.001 | -0.001 | 0.003 | 0.005 | ... | -0.001 | 0.000 | 0.001 | -0.001 | -0.003 | -0.002 | 0.002 | 0.001 | 0.002 | 0.000 |
| Actinomyces | -0.000 | 0.002 | 0.001 | -0.115 | 0.002 | -0.002 | 0.001 | 0.002 | 0.002 | 0.005 | ... | -0.002 | 0.002 | -0.001 | -0.002 | -0.005 | 0.003 | -0.000 | 0.000 | -0.002 | -0.002 |
| Flavobacterium | -0.002 | -0.000 | 0.002 | -0.002 | -0.049 | -0.002 | 0.001 | -0.003 | -0.003 | 0.001 | ... | -0.001 | 0.002 | -0.001 | 0.002 | -0.001 | -0.001 | 0.002 | -0.001 | 0.001 | 0.001 |
| Prevotella | -0.001 | -0.000 | -0.001 | -0.002 | -0.002 | -0.047 | 0.002 | -0.002 | -0.002 | 0.004 | ... | -0.001 | 0.002 | 0.002 | -0.001 | 0.001 | 0.001 | -0.002 | 0.002 | 0.000 | 0.001 |
| Porphyromonas | -0.000 | 0.003 | -0.002 | -0.001 | 0.001 | 0.001 | -0.017 | -0.001 | -0.001 | -0.000 | ... | -0.001 | -0.002 | 0.000 | 0.000 | 0.004 | 0.000 | -0.002 | -0.001 | -0.002 | 0.000 |
| Parvimonas | 0.002 | -0.001 | -0.002 | -0.001 | 0.002 | -0.000 | -0.002 | 0.263 | -0.001 | -0.001 | ... | -0.000 | -0.000 | -0.000 | 0.001 | -0.000 | -0.001 | -0.001 | 0.002 | -0.001 | 0.001 |
| Fusobacterium | -0.000 | -0.000 | -0.001 | -0.000 | 0.000 | 0.001 | -0.001 | 0.000 | 0.265 | -0.001 | ... | -0.002 | -0.002 | 0.000 | 0.002 | -0.000 | -0.001 | 0.002 | 0.001 | -0.001 | 0.002 |
| Propionibacterium | 0.001 | 0.001 | -0.001 | 0.002 | -0.001 | 0.003 | 0.000 | -0.006 | -0.004 | -2.003 | ... | 0.000 | -0.000 | -0.001 | -0.001 | 0.008 | -0.002 | -0.000 | -0.000 | -0.002 | 0.001 |
| Gemella | 0.002 | -0.002 | 0.000 | -0.001 | -0.002 | 0.000 | 0.001 | -0.003 | -0.003 | 0.002 | ... | -0.001 | -0.001 | 0.002 | -0.002 | 0.001 | -0.001 | 0.000 | 0.001 | 0.000 | 0.002 |
| Rothia | 0.001 | 0.001 | 0.000 | -0.002 | -0.002 | 0.001 | 0.001 | -0.008 | -0.008 | -0.011 | ... | 0.001 | -0.002 | -0.006 | -0.003 | 0.009 | -0.002 | -0.002 | 0.001 | -0.001 | 0.002 |
| Granulicatella | -0.002 | -0.001 | 0.001 | -0.000 | -0.002 | 0.001 | 0.001 | -0.000 | -0.001 | 0.001 | ... | -0.001 | -0.002 | 0.001 | -0.001 | -0.001 | -0.001 | -0.002 | 0.001 | -0.001 | -0.000 |
| Neisseria | -0.000 | 0.002 | 0.002 | -0.002 | 0.001 | 0.002 | 0.001 | -0.004 | -0.000 | -0.002 | ... | -0.001 | 0.002 | 0.002 | -0.002 | 0.004 | 0.001 | 0.001 | 0.002 | -0.000 | -0.001 |
| Lactobacillus | -0.002 | 0.000 | 0.001 | 0.001 | -0.001 | 0.000 | -0.001 | 0.002 | -0.001 | 0.002 | ... | -0.001 | 0.001 | 0.001 | 0.000 | -0.001 | -0.001 | -0.001 | 0.002 | 0.000 | -0.000 |
| Megasphaera | -0.000 | 0.000 | -0.251 | 0.002 | 0.001 | 0.002 | 0.001 | -0.002 | -0.004 | -0.004 | ... | 0.000 | 0.000 | 0.002 | -0.001 | 0.001 | 0.001 | 0.000 | -0.001 | 0.001 | -0.002 |
| Catonella | -0.002 | -0.001 | -0.002 | 0.002 | 0.002 | -0.002 | -0.002 | -0.002 | 0.003 | 0.002 | ... | -0.001 | -0.001 | 0.002 | 0.000 | -0.003 | 0.001 | 0.001 | -0.002 | -0.001 | -0.002 |
| Atopobium | -0.002 | 0.002 | 0.002 | -0.002 | -0.003 | 0.000 | -0.000 | -0.001 | -0.003 | -0.001 | ... | -0.002 | -0.001 | 0.001 | 0.001 | 0.004 | 0.001 | -0.003 | -0.001 | -0.002 | -0.002 |
| Campylobacter | -0.002 | 0.000 | 0.002 | 0.002 | -0.001 | -0.002 | 0.002 | -0.003 | 0.001 | -0.000 | ... | 0.000 | -0.001 | -0.002 | -0.003 | -0.001 | -0.001 | 0.000 | 0.003 | -0.002 | -0.001 |
| Capnocytophaga | 0.002 | 0.002 | -0.002 | 0.000 | 0.036 | 0.002 | -0.000 | 0.002 | 0.002 | 0.000 | ... | -0.001 | -0.002 | 0.000 | -0.000 | -0.003 | -0.002 | 0.003 | 0.001 | 0.002 | -0.002 |
| Solobacterium | 0.000 | -0.001 | -0.002 | 0.001 | -0.001 | -0.001 | -0.001 | -0.002 | 0.002 | 0.001 | ... | -0.002 | -0.002 | -0.001 | 0.000 | -0.002 | -0.000 | -0.001 | -0.002 | 0.002 | -0.002 |
| Moryella | -0.000 | -0.002 | 0.002 | -0.002 | -0.001 | 0.001 | -0.001 | 0.004 | 0.007 | 0.006 | ... | 0.002 | -0.001 | 0.003 | -0.000 | -0.005 | -0.002 | -0.001 | 0.001 | 0.002 | -0.001 |
| TM7_genera_incertae_sedis | 0.000 | -0.001 | 0.002 | -0.002 | 0.000 | 0.002 | 0.002 | -0.001 | -0.002 | -0.002 | ... | -0.002 | 0.000 | 0.000 | -0.002 | 0.005 | -0.002 | 0.001 | -0.002 | -0.002 | -0.002 |
| Staphylococcus | -0.002 | 0.002 | -0.001 | 0.002 | 0.001 | -0.002 | -0.000 | -0.003 | 0.002 | 0.001 | ... | 0.002 | 0.003 | -0.001 | -0.001 | 0.000 | 0.000 | -0.001 | 0.002 | 0.000 | -0.002 |
| Filifactor | -0.002 | 0.002 | 0.002 | -0.002 | 0.000 | -0.001 | -0.002 | 0.002 | 0.003 | 0.005 | ... | -0.000 | -0.001 | 0.001 | -0.002 | -0.002 | 0.000 | -0.001 | 0.000 | -0.001 | -0.001 |
| Oribacterium | 0.000 | -0.001 | 0.002 | 0.000 | -0.001 | -0.003 | 0.003 | 0.004 | 0.009 | 0.009 | ... | -0.002 | 0.001 | 0.001 | 0.003 | -0.005 | 0.003 | 0.003 | -0.000 | -0.000 | -0.000 |
| Burkholderia | 0.000 | 0.001 | 0.002 | 0.000 | -0.001 | -0.001 | 0.001 | -0.001 | -0.001 | -0.000 | ... | 0.002 | -0.000 | 0.001 | 0.000 | 0.004 | 0.000 | -0.002 | -0.002 | 0.002 | -0.000 |
| Sneathia | -0.001 | 0.000 | -0.003 | 0.000 | -0.001 | -0.001 | 0.000 | -0.002 | -0.002 | -0.000 | ... | -0.001 | 0.002 | 0.000 | -0.001 | 0.003 | -0.002 | 0.002 | -0.002 | 0.001 | 0.001 |
| Treponema | 0.002 | 0.001 | -0.002 | 0.001 | -0.001 | -0.000 | -0.002 | -0.002 | 0.003 | 0.002 | ... | -0.001 | 0.002 | 0.002 | 0.002 | -0.001 | 0.000 | -0.001 | 0.001 | -0.001 | 0.002 |
| Moraxella | 0.001 | -0.002 | -0.001 | 0.000 | -0.002 | 0.002 | 0.002 | 0.000 | 0.002 | -0.002 | ... | -0.001 | -0.000 | 0.002 | 0.001 | -0.000 | -0.000 | -0.003 | 0.000 | -0.000 | 0.002 |
| Haemophilus | 0.001 | 0.002 | -0.002 | -0.001 | -0.001 | 0.002 | -0.000 | -0.001 | 0.002 | -0.000 | ... | 0.001 | -0.000 | -0.000 | 0.001 | -0.001 | -0.001 | 0.002 | 0.002 | -0.002 | -0.002 |
| Selenomonas | 0.001 | 0.002 | 0.107 | -0.001 | 0.000 | -0.001 | -0.001 | 0.001 | 0.001 | 0.005 | ... | -0.001 | -0.002 | -0.001 | 0.003 | -0.002 | 0.001 | -0.002 | 0.002 | 0.002 | 0.002 |
| Corynebacterium | -0.001 | 0.000 | -0.003 | 0.002 | 0.002 | 0.001 | 0.001 | -0.002 | -0.000 | -0.005 | ... | -0.002 | -0.002 | 0.002 | 0.000 | 0.001 | -0.002 | 0.000 | -0.000 | -0.002 | 0.000 |
| Rhizobium | -0.000 | -0.001 | -0.001 | -0.000 | -0.002 | -0.001 | 0.001 | -0.002 | 0.002 | 0.001 | ... | -0.002 | -0.001 | 0.002 | -0.002 | 0.001 | -0.002 | 0.001 | 0.002 | -0.001 | 0.002 |
| Bradyrhizobium | -0.000 | -0.002 | -0.001 | 0.001 | 0.002 | -0.002 | 0.002 | -0.002 | -0.003 | -0.002 | ... | -0.003 | 0.001 | 0.000 | -0.001 | 0.004 | -0.000 | -0.002 | -0.001 | 0.000 | 0.001 |
| Methylobacterium | 0.002 | 0.001 | -0.001 | -0.000 | -0.001 | -0.002 | 0.001 | -0.003 | 0.001 | 0.001 | ... | 0.672 | 0.000 | 0.002 | -0.000 | 0.000 | 0.002 | 0.002 | -0.001 | -0.001 | -0.001 |
| OD1_genera_incertae_sedis | 0.001 | -0.002 | 0.000 | -0.002 | 0.002 | 0.001 | 0.001 | -0.000 | -0.002 | 0.002 | ... | 0.002 | -0.299 | 0.001 | 0.000 | -0.001 | 0.003 | -0.000 | 0.001 | 0.002 | 0.002 |
| Finegoldia | 0.002 | -0.001 | 0.001 | -0.001 | 0.002 | -0.002 | 0.001 | 0.152 | 0.001 | 0.001 | ... | 0.002 | -0.001 | -0.001 | 0.002 | 0.001 | 0.001 | -0.002 | -0.002 | -0.002 | -0.000 |
| Microbacterium | 0.001 | -0.000 | 0.002 | -0.002 | -0.001 | -0.002 | 0.002 | -0.003 | 0.001 | -0.000 | ... | -0.000 | 0.001 | -0.416 | 0.002 | -0.001 | 0.001 | 0.001 | 0.002 | -0.001 | -0.002 |
| Sphingomonas | 0.001 | 0.002 | 0.002 | -0.002 | -0.001 | -0.002 | 0.000 | 0.000 | -0.002 | 0.001 | ... | 0.000 | 0.000 | 0.001 | -0.376 | 0.004 | -0.003 | -0.003 | 0.001 | 0.000 | -0.000 |
| Chryseobacterium | -0.002 | -0.000 | 0.003 | 0.000 | 0.269 | -0.002 | -0.000 | 0.001 | 0.000 | -0.001 | ... | 0.001 | 0.001 | 0.000 | -0.000 | 0.002 | -0.001 | 0.000 | 0.001 | 0.001 | -0.000 |
| Bacteroides | 0.001 | -0.002 | 0.001 | 0.001 | 0.001 | -0.000 | 0.002 | -0.001 | -0.001 | -0.003 | ... | -0.001 | 0.001 | 0.001 | 0.001 | 1.461 | -0.002 | -0.003 | -0.002 | -0.001 | -0.001 |
| Bdellovibrio | 0.000 | 0.002 | -0.003 | -0.001 | -0.001 | 0.000 | -0.001 | 0.001 | 0.003 | 0.001 | ... | -0.002 | 0.002 | 0.000 | 0.001 | 0.001 | -0.146 | 0.003 | -0.001 | 0.000 | 0.000 |
| Streptophyta | -0.002 | -0.001 | -0.001 | 0.002 | 0.001 | -0.001 | 0.003 | 0.001 | -0.002 | 0.000 | ... | -0.001 | -0.001 | -0.001 | 0.002 | -0.002 | -0.000 | -0.147 | -0.002 | -0.001 | 0.002 |
| Lachnospiracea_incertae_sedis | 0.000 | 0.001 | -0.000 | -0.001 | -0.001 | 0.002 | -0.001 | 0.001 | -0.000 | -0.001 | ... | 0.002 | 0.002 | -0.001 | 0.001 | 0.002 | 0.001 | 0.002 | -0.002 | -0.002 | -0.000 |
| Paracoccus | 0.001 | 0.002 | 0.001 | 0.001 | 0.000 | 0.001 | -0.002 | -0.001 | -0.001 | 0.002 | ... | 0.002 | -0.002 | -0.001 | -0.001 | 0.001 | -0.001 | -0.001 | 0.252 | 0.001 | -0.001 |
| Fastidiosipila | 0.003 | 0.000 | 0.000 | -0.002 | -0.001 | -0.002 | 0.000 | -0.001 | 0.002 | -0.001 | ... | -0.001 | -0.002 | 0.001 | 0.001 | -0.001 | 0.000 | -0.001 | 0.001 | 0.087 | -0.002 |
| Pseudonocardia | 0.000 | 0.001 | -0.001 | -0.002 | -0.002 | -0.001 | -0.001 | 0.001 | 0.000 | 0.004 | ... | 0.002 | -0.002 | -0.000 | -0.003 | 0.000 | -0.000 | 0.002 | -0.000 | 0.002 | 0.071 |
48 rows × 40 columns
9. Taxa selection performance¶
If we know the true disease path, we can calculate the taxa selection performance by deepbiome_taxa_selection_performance funciton. First, we prepared the true weight list based on the true disease path. For each fold, we prepared 4 weights from the 5 layers ([genus to family, family to order, order to Class, class to phylum]). An example of the list of the true weights from each fold is as follow:
[44]:
true_tree_weight_list = np.load(resource_filename('deepbiome', 'tests/data/true_weight_list.npy'), allow_pickle=True)
log.info(true_tree_weight_list.shape)
[root |INFO|<ipython-input-44-7f16305fbcb7>:2] (5, 4)
The first weight between the genus and family layers for first epoch has the shape below:
[45]:
log.info(true_tree_weight_list[0][0].shape)
[root |INFO|<ipython-input-45-7f1406e7d9a7>:1] (48, 40)
We will calculate the taxa selection performance of the trained weight below:
[46]:
trained_weight_path_list = ['%s/weight_%d.h5' % (path_info['model_info']['model_dir'], i) for i in range(5)]
trained_weight_path_list
[46]:
['./example_result//weight_0.h5',
'./example_result//weight_1.h5',
'./example_result//weight_2.h5',
'./example_result//weight_3.h5',
'./example_result//weight_4.h5']
This is the summary of the taxa selection accuracy of trained weights from each fold.
[47]:
summary = deepbiome.deepbiome_taxa_selection_performance(log, network_info, path_info, num_classes=1,
true_tree_weight_list=true_tree_weight_list,
trained_weight_path_list = trained_weight_path_list)
[root |INFO|readers.py:58] -----------------------------------------------------------------------
[root |INFO|readers.py:59] Construct Dataset
[root |INFO|readers.py:60] -----------------------------------------------------------------------
[root |INFO|readers.py:61] Load data
[root |INFO|build_network.py:87] Load trained model weight at ./example_result//weight_0.h5
[root |INFO|build_network.py:87] Load trained model weight at ./example_result//weight_1.h5
[root |INFO|build_network.py:87] Load trained model weight at ./example_result//weight_2.h5
[root |INFO|build_network.py:87] Load trained model weight at ./example_result//weight_3.h5
[root |INFO|build_network.py:87] Load trained model weight at ./example_result//weight_4.h5
[48]:
summary
[48]:
| Model | PhyloTree | No. true taxa | No. total taxa | Sensitivity_mean | Sensitivity_std | Specificity_mean | Specificity_std | Gmeasure_mean | Gmeasure_std | Accuracy_mean | Accuracy_std | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 0 | ./example_result/ | Genus | 31 | 48 | 0.948 | 0.044 | 0.983 | 0.014 | 0.965 | 0.021 | 0.982 | 0.013 |
| 1 | Family | 23 | 40 | 0.991 | 0.017 | 0.973 | 0.011 | 0.982 | 0.008 | 0.973 | 0.010 | |
| 2 | Order | 9 | 23 | 0.978 | 0.044 | 0.953 | 0.015 | 0.965 | 0.021 | 0.954 | 0.015 | |
| 3 | Class | 7 | 17 | 0.971 | 0.057 | 0.925 | 0.018 | 0.948 | 0.037 | 0.927 | 0.020 |
10. Draw phylogenetic tree¶
The deepbiome_get_trained_weight function draw phylogenetic tree for the DeepBiome network. This function implemented ete3 library (link TBD).
If we turn on the tree_weight_on option, the function shows the amount and direction of the tree weights for each edge by circle size and color. For this option, we have to feed a list of pandas dataframe of the tree weights obtained from deepbiome_get_trained_weight function to tree_weight argument. In this example, we used the 5 lists of weights from 5-fold cross validation obtained from deepbiome_get_trained_weight function. For each layer, we first averaged 5 weights from 5
fold CV.
[49]:
tot_trained_weight_list = []
for fold in range(5):
weight_path = '%s/%s' % (path_info['model_info']['model_dir'], 'weight_%d.h5' % fold)
trained_weight_list = deepbiome.deepbiome_get_trained_weight(log, network_info, path_info, num_classes=1,
weight_path=weight_path, verbose=False)
tot_trained_weight_list.append(trained_weight_list)
trained_weight_list = []
for i in range(len(tot_trained_weight_list[0])):
level_weights = tot_trained_weight_list[0][i]
for j in range(1,len(tot_trained_weight_list)):
level_weights +=tot_trained_weight_list[j][i]
level_weights /= len(tot_trained_weight_list)
trained_weight_list.append(level_weights)
Now, we can draw the phylogenetic tree with trained weights. For using the trained weights, ``tree_level_list``, the list of names for each level, should be same as the name of each level in the trained weight list. We used file_name='%%inline' option to show the tree in the jupyter notebook. To specify the background color for each phylum, we turned on the phylum_background_color_on option and used phylum_color list.
[50]:
phylum_color = ['lightsteelblue',
'moccasin',
'darkseagreen',
'khaki',
'mediumturquoise',
'lightblue',
'thistle',
'tan',
'mistyrose',
]
img = deepbiome.deepbiome_draw_phylogenetic_tree(log, network_info, path_info, num_classes=1,
file_name='%%inline', img_w=500, branch_vertical_margin=20,
arc_start=0, arc_span=360,
node_name_on=False, name_fsize=10,
tree_weight_on=True, tree_weight=trained_weight_list,
tree_level_list = ['Genus', 'Family', 'Order', 'Class', 'Phylum', 'Disease'],
weight_opacity=0.8, weight_max_radios=10,
phylum_background_color_on=True, phylum_color=phylum_color, phylum_color_legend=True,
verbose=False)
img
[50]:
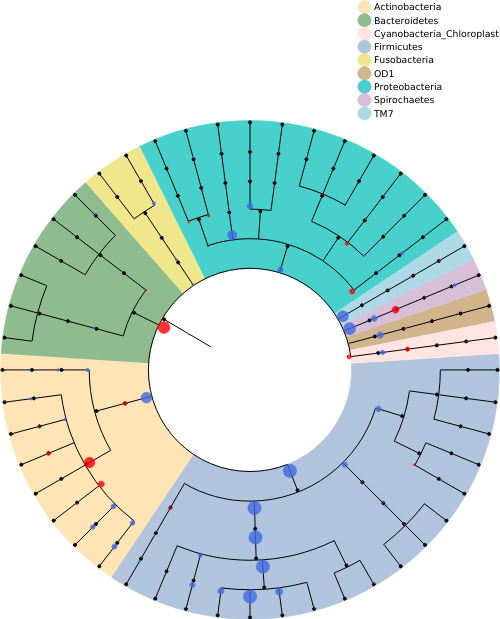
We can show the name of each leaf node in the phylogenetic tree by turn on the node_name_on option.
[51]:
img = deepbiome.deepbiome_draw_phylogenetic_tree(log, network_info, path_info, num_classes=1,
file_name='%%inline', img_w=500,
branch_vertical_margin=20, arc_start=0, arc_span=360,
node_name_on=True, name_fsize=20,
tree_weight_on=True, tree_weight=trained_weight_list,
tree_level_list = ['Genus', 'Family', 'Order', 'Class', 'Phylum', 'Disease'],
weight_opacity=0.8, weight_max_radios=20,
phylum_background_color_on=True, phylum_color=phylum_color, phylum_color_legend=True,
verbose=False)
img
[51]:
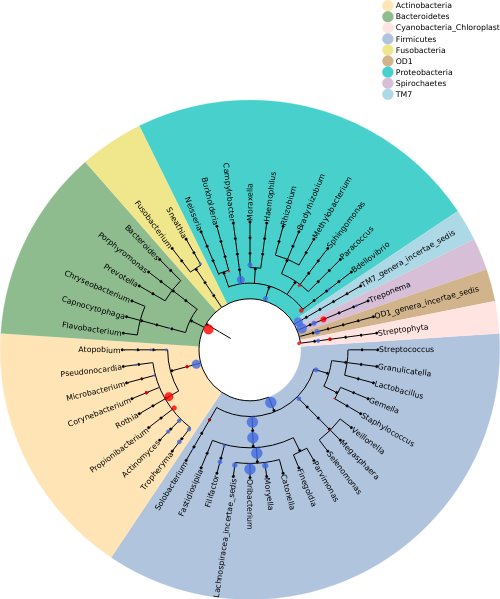
If we want to save the figure as png or jpg, we can change file_name option.
[52]:
img = deepbiome.deepbiome_draw_phylogenetic_tree(log, network_info, path_info, num_classes=1,
file_name='tree.png', img_w=500,
branch_vertical_margin=20, arc_start=0, arc_span=360,
node_name_on=True, name_fsize=20,
tree_weight_on=True, tree_weight=trained_weight_list,
tree_level_list = ['Genus', 'Family', 'Order', 'Class', 'Phylum', 'Disease'],
weight_opacity=0.8, weight_max_radios=20,
phylum_background_color_on=True, phylum_color=phylum_color, phylum_color_legend=True,
verbose=False)
We can check that the tree image saved in tree.png.
[53]:
img = plt.imread('tree.png')
plt.imshow(img)
plt.show()
11. Using covariates¶
We can use covariates for training the DeepBiome network.
Below is an example of the input files for the continuous covariates.
[54]:
pd.read_csv(resource_filename('deepbiome', 'tests/data/ages.csv'))
[54]:
| ages | |
|---|---|
| 0 | 59.000 |
| 1 | 45.000 |
| 2 | 42.000 |
| 3 | 50.000 |
| 4 | 69.000 |
| 5 | 37.000 |
| 6 | 66.000 |
| 7 | 75.000 |
| 8 | 32.000 |
| 9 | 72.000 |
| 10 | 65.000 |
| 11 | 58.000 |
| 12 | 41.000 |
| 13 | 49.000 |
| 14 | 49.000 |
| 15 | 70.000 |
| 16 | 64.000 |
| 17 | 33.000 |
| 18 | 74.000 |
| 19 | 56.000 |
| 20 | 37.000 |
| 21 | 38.000 |
| 22 | 51.000 |
| 23 | 34.000 |
| 24 | 46.000 |
| 25 | 44.000 |
| 26 | 60.000 |
| 27 | 46.000 |
| 28 | 37.000 |
| 29 | 58.000 |
| ... | ... |
| 970 | 38.000 |
| 971 | 78.000 |
| 972 | 32.000 |
| 973 | 62.000 |
| 974 | 47.000 |
| 975 | 47.000 |
| 976 | 65.000 |
| 977 | 68.000 |
| 978 | 45.000 |
| 979 | 43.000 |
| 980 | 31.000 |
| 981 | 45.000 |
| 982 | 64.000 |
| 983 | 36.000 |
| 984 | 36.000 |
| 985 | 59.000 |
| 986 | 78.000 |
| 987 | 49.000 |
| 988 | 33.000 |
| 989 | 60.000 |
| 990 | 48.000 |
| 991 | 75.000 |
| 992 | 38.000 |
| 993 | 39.000 |
| 994 | 49.000 |
| 995 | 71.000 |
| 996 | 34.000 |
| 997 | 48.000 |
| 998 | 38.000 |
| 999 | 52.000 |
1000 rows × 1 columns
Below is an example of the input files for the categorical covariates. We assume that the file contains all of the catetogories for one-hot encoding.
[55]:
pd.read_csv(resource_filename('deepbiome', 'tests/data/gender.csv'))
[55]:
| gender | |
|---|---|
| 0 | 1 |
| 1 | 0 |
| 2 | 1 |
| 3 | 1 |
| 4 | 1 |
| 5 | 0 |
| 6 | 1 |
| 7 | 1 |
| 8 | 0 |
| 9 | 0 |
| 10 | 0 |
| 11 | 1 |
| 12 | 0 |
| 13 | 1 |
| 14 | 1 |
| 15 | 1 |
| 16 | 0 |
| 17 | 0 |
| 18 | 1 |
| 19 | 1 |
| 20 | 0 |
| 21 | 0 |
| 22 | 1 |
| 23 | 0 |
| 24 | 1 |
| 25 | 0 |
| 26 | 0 |
| 27 | 0 |
| 28 | 1 |
| 29 | 1 |
| ... | ... |
| 970 | 1 |
| 971 | 1 |
| 972 | 0 |
| 973 | 1 |
| 974 | 0 |
| 975 | 1 |
| 976 | 0 |
| 977 | 1 |
| 978 | 1 |
| 979 | 1 |
| 980 | 1 |
| 981 | 1 |
| 982 | 0 |
| 983 | 0 |
| 984 | 0 |
| 985 | 1 |
| 986 | 0 |
| 987 | 0 |
| 988 | 1 |
| 989 | 0 |
| 990 | 0 |
| 991 | 1 |
| 992 | 1 |
| 993 | 1 |
| 994 | 0 |
| 995 | 0 |
| 996 | 0 |
| 997 | 1 |
| 998 | 0 |
| 999 | 1 |
1000 rows × 1 columns
[56]:
pd.read_csv(resource_filename('deepbiome', 'tests/data/ethnicity.csv'))
[56]:
| ethnicity | |
|---|---|
| 0 | 4 |
| 1 | 0 |
| 2 | 0 |
| 3 | 4 |
| 4 | 4 |
| 5 | 2 |
| 6 | 3 |
| 7 | 0 |
| 8 | 2 |
| 9 | 5 |
| 10 | 2 |
| 11 | 3 |
| 12 | 5 |
| 13 | 3 |
| 14 | 0 |
| 15 | 0 |
| 16 | 4 |
| 17 | 5 |
| 18 | 1 |
| 19 | 1 |
| 20 | 4 |
| 21 | 4 |
| 22 | 1 |
| 23 | 0 |
| 24 | 1 |
| 25 | 3 |
| 26 | 1 |
| 27 | 4 |
| 28 | 4 |
| 29 | 3 |
| ... | ... |
| 970 | 1 |
| 971 | 0 |
| 972 | 0 |
| 973 | 5 |
| 974 | 3 |
| 975 | 2 |
| 976 | 1 |
| 977 | 5 |
| 978 | 4 |
| 979 | 1 |
| 980 | 0 |
| 981 | 3 |
| 982 | 3 |
| 983 | 5 |
| 984 | 5 |
| 985 | 5 |
| 986 | 3 |
| 987 | 1 |
| 988 | 2 |
| 989 | 2 |
| 990 | 0 |
| 991 | 5 |
| 992 | 3 |
| 993 | 5 |
| 994 | 5 |
| 995 | 2 |
| 996 | 2 |
| 997 | 1 |
| 998 | 0 |
| 999 | 0 |
1000 rows × 1 columns
For using the covariates, we have to set the covariates_info field in the configuration for the path informations. If we don’t specify the covariates_info, than the DeepBiome will train the model without any covariates.
[57]:
network_info_with_covariates = {
'architecture_info': {
'batch_normalization': 'False',
'drop_out': '0',
'weight_initial': 'glorot_uniform',
'weight_l1_penalty':'0.',
'weight_decay': 'phylogenetic_tree',
},
'model_info': {
'lr': '0.01',
'decay': '0.001',
'loss': 'binary_crossentropy',
'metrics': 'binary_accuracy, sensitivity, specificity, gmeasure',
'taxa_selection_metrics': 'sensitivity, specificity, gmeasure, accuracy',
'network_class': 'DeepBiomeNetwork',
'optimizer': 'adam',
'reader_class': 'MicroBiomeClassificationReader',
'normalizer': 'normalize_minmax',
},
'training_info': {
'epochs': '10',
'batch_size': '50',
'callbacks': 'ModelCheckpoint',
'monitor': 'val_loss',
'mode' : 'min',
'min_delta': '1e-7',
},
'validation_info': {
'batch_size': 'None',
'validation_size': '0.2'
},
'test_info': {
'batch_size': 'None'
}
}
[58]:
path_info_with_covariates = {
'data_info': {
'data_path': resource_filename('deepbiome', 'tests/data'),
'idx_path': resource_filename('deepbiome', 'tests/data/onefile_idx.csv'),
'tree_info_path': resource_filename('deepbiome', 'tests/data/genus48_dic.csv'),
'x_path': 'onefile_x.csv',
'y_path': 'classification_y.csv'
},
'model_info': {
'evaluation': 'eval.npy',
'history': 'hist.json',
'model_dir': './example_result/',
'weight': 'weight.h5'
},
'covariates_info':{
'continuous_variables':','.join([resource_filename('deepbiome', 'tests/data/ages.csv'),
resource_filename('deepbiome', 'tests/data/weights.csv')]),
'categorical_variables':','.join([resource_filename('deepbiome', 'tests/data/gender.csv'),
resource_filename('deepbiome', 'tests/data/ethnicity.csv')]),
}
}
We can use the same deepbiome_train function when we train the network with covariates.
[59]:
test_evaluation_with_cov, train_evaluation_with_cov, network_with_cov = deepbiome.deepbiome_train(log, network_info_with_covariates,
path_info_with_covariates,
number_of_fold=None)
[root |INFO|deepbiome.py:115] -----------------------------------------------------------------
[root |INFO|deepbiome.py:153] -------1 simulation start!----------------------------------
[root |INFO|readers.py:58] -----------------------------------------------------------------------
[root |INFO|readers.py:59] Construct Dataset
[root |INFO|readers.py:60] -----------------------------------------------------------------------
[root |INFO|readers.py:61] Load data
[root |INFO|deepbiome.py:164] -----------------------------------------------------------------
[root |INFO|deepbiome.py:165] Build network for 1 simulation
[root |INFO|build_network.py:521] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:522] Read phylogenetic tree information from /DATA/home/muha/github_repos/deepbiome/deepbiome/tests/data/genus48_dic.csv
[root |INFO|build_network.py:528] Phylogenetic tree level list: ['Genus', 'Family', 'Order', 'Class', 'Phylum']
[root |INFO|build_network.py:529] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:537] Genus: 48
[root |INFO|build_network.py:537] Family: 40
[root |INFO|build_network.py:537] Order: 23
[root |INFO|build_network.py:537] Class: 17
[root |INFO|build_network.py:537] Phylum: 9
[root |INFO|build_network.py:543] Phylum_with_covariates: 17
[root |INFO|build_network.py:546] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:547] Phylogenetic_tree_dict info: ['Genus', 'Order', 'Class', 'Family', 'Phylum_with_covariates', 'Phylum', 'Number']
[root |INFO|build_network.py:548] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:558] Build edge weights between [ Genus, Family]
[root |INFO|build_network.py:558] Build edge weights between [Family, Order]
[root |INFO|build_network.py:558] Build edge weights between [ Order, Class]
[root |INFO|build_network.py:558] Build edge weights between [ Class, Phylum]
[root |INFO|build_network.py:571] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:586] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:587] Build network based on phylogenetic tree information
[root |INFO|build_network.py:588] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:670] ------------------------------------------------------------------------------------------
Model: "model_4"
__________________________________________________________________________________________________
Layer (type) Output Shape Param # Connected to
==================================================================================================
input (InputLayer) (None, 48) 0
__________________________________________________________________________________________________
l1_dense (Dense_with_tree) (None, 40) 1960 input[0][0]
__________________________________________________________________________________________________
l1_activation (Activation) (None, 40) 0 l1_dense[0][0]
__________________________________________________________________________________________________
l2_dense (Dense_with_tree) (None, 23) 943 l1_activation[0][0]
__________________________________________________________________________________________________
l2_activation (Activation) (None, 23) 0 l2_dense[0][0]
__________________________________________________________________________________________________
l3_dense (Dense_with_tree) (None, 17) 408 l2_activation[0][0]
__________________________________________________________________________________________________
l3_activation (Activation) (None, 17) 0 l3_dense[0][0]
__________________________________________________________________________________________________
l4_dense (Dense_with_tree) (None, 9) 162 l3_activation[0][0]
__________________________________________________________________________________________________
l4_activation (Activation) (None, 9) 0 l4_dense[0][0]
__________________________________________________________________________________________________
covariates_input (InputLayer) (None, 8) 0
__________________________________________________________________________________________________
biome_covariates_concat (Concat (None, 17) 0 l4_activation[0][0]
covariates_input[0][0]
__________________________________________________________________________________________________
last_dense_h (Dense) (None, 1) 18 biome_covariates_concat[0][0]
__________________________________________________________________________________________________
p_hat (Activation) (None, 1) 0 last_dense_h[0][0]
==================================================================================================
Total params: 3,491
Trainable params: 3,491
Non-trainable params: 0
__________________________________________________________________________________________________
[root |INFO|build_network.py:61] Build Network
[root |INFO|build_network.py:62] Optimizer = adam
[root |INFO|build_network.py:63] Loss = binary_crossentropy
[root |INFO|build_network.py:64] Metrics = binary_accuracy, sensitivity, specificity, gmeasure
[root |INFO|deepbiome.py:176] -----------------------------------------------------------------
[root |INFO|deepbiome.py:177] 1 fold computing start!----------------------------------
[root |INFO|build_network.py:137] Training start!
Train on 640 samples, validate on 160 samples
Epoch 1/10
640/640 [==============================] - 1s 1ms/step - loss: 11.0308 - binary_accuracy: 0.3156 - sensitivity: 0.0000e+00 - specificity: 1.0000 - gmeasure: 0.0000e+00 - val_loss: 11.1819 - val_binary_accuracy: 0.3063 - val_sensitivity: 0.0000e+00 - val_specificity: 1.0000 - val_gmeasure: 0.0000e+00
Epoch 2/10
640/640 [==============================] - 0s 101us/step - loss: 11.0308 - binary_accuracy: 0.3156 - sensitivity: 0.0000e+00 - specificity: 1.0000 - gmeasure: 0.0000e+00 - val_loss: 11.1819 - val_binary_accuracy: 0.3063 - val_sensitivity: 0.0000e+00 - val_specificity: 1.0000 - val_gmeasure: 0.0000e+00
Epoch 3/10
640/640 [==============================] - 0s 101us/step - loss: 11.0308 - binary_accuracy: 0.3156 - sensitivity: 0.0000e+00 - specificity: 1.0000 - gmeasure: 0.0000e+00 - val_loss: 11.1819 - val_binary_accuracy: 0.3063 - val_sensitivity: 0.0000e+00 - val_specificity: 1.0000 - val_gmeasure: 0.0000e+00
Epoch 4/10
640/640 [==============================] - 0s 105us/step - loss: 11.0308 - binary_accuracy: 0.3156 - sensitivity: 0.0000e+00 - specificity: 1.0000 - gmeasure: 0.0000e+00 - val_loss: 11.1819 - val_binary_accuracy: 0.3063 - val_sensitivity: 0.0000e+00 - val_specificity: 1.0000 - val_gmeasure: 0.0000e+00
Epoch 5/10
640/640 [==============================] - 0s 130us/step - loss: 11.0308 - binary_accuracy: 0.3156 - sensitivity: 0.0000e+00 - specificity: 1.0000 - gmeasure: 0.0000e+00 - val_loss: 11.1819 - val_binary_accuracy: 0.3063 - val_sensitivity: 0.0000e+00 - val_specificity: 1.0000 - val_gmeasure: 0.0000e+00
Epoch 6/10
640/640 [==============================] - 0s 123us/step - loss: 11.0308 - binary_accuracy: 0.3156 - sensitivity: 0.0000e+00 - specificity: 1.0000 - gmeasure: 0.0000e+00 - val_loss: 11.1819 - val_binary_accuracy: 0.3063 - val_sensitivity: 0.0000e+00 - val_specificity: 1.0000 - val_gmeasure: 0.0000e+00
Epoch 7/10
640/640 [==============================] - 0s 120us/step - loss: 11.0308 - binary_accuracy: 0.3156 - sensitivity: 0.0000e+00 - specificity: 1.0000 - gmeasure: 0.0000e+00 - val_loss: 11.1819 - val_binary_accuracy: 0.3063 - val_sensitivity: 0.0000e+00 - val_specificity: 1.0000 - val_gmeasure: 0.0000e+00
Epoch 8/10
640/640 [==============================] - 0s 120us/step - loss: 11.0308 - binary_accuracy: 0.3156 - sensitivity: 0.0000e+00 - specificity: 1.0000 - gmeasure: 0.0000e+00 - val_loss: 11.1819 - val_binary_accuracy: 0.3063 - val_sensitivity: 0.0000e+00 - val_specificity: 1.0000 - val_gmeasure: 0.0000e+00
Epoch 9/10
640/640 [==============================] - 0s 114us/step - loss: 11.0308 - binary_accuracy: 0.3156 - sensitivity: 0.0000e+00 - specificity: 1.0000 - gmeasure: 0.0000e+00 - val_loss: 11.1819 - val_binary_accuracy: 0.3063 - val_sensitivity: 0.0000e+00 - val_specificity: 1.0000 - val_gmeasure: 0.0000e+00
Epoch 10/10
640/640 [==============================] - 0s 103us/step - loss: 11.0308 - binary_accuracy: 0.3156 - sensitivity: 0.0000e+00 - specificity: 1.0000 - gmeasure: 0.0000e+00 - val_loss: 11.1819 - val_binary_accuracy: 0.3063 - val_sensitivity: 0.0000e+00 - val_specificity: 1.0000 - val_gmeasure: 0.0000e+00
[root |INFO|build_network.py:87] Load trained model weight at ./example_result/weight_0.h5
[root |INFO|build_network.py:147] Training end with time 3.0836904048919678!
[root |INFO|build_network.py:83] Saved trained model weight at ./example_result/weight_0.h5
[root |DEBUG|deepbiome.py:185] Save weight at ./example_result/weight_0.h5
[root |DEBUG|deepbiome.py:188] Save history at ./example_result/hist_0.json
[root |INFO|build_network.py:173] Evaluation start!
800/800 [==============================] - 0s 3us/step
[root |INFO|build_network.py:178] Evaluation end with time 0.010069608688354492!
[root |INFO|build_network.py:179] Evaluation: [11.061043739318848, 0.3137499988079071, 0.0, 1.0, 0.0]
[root |INFO|build_network.py:173] Evaluation start!
200/200 [==============================] - 0s 11us/step
[root |INFO|build_network.py:178] Evaluation end with time 0.008058786392211914!
[root |INFO|build_network.py:179] Evaluation: [11.121485710144043, 0.3100000023841858, 0.0, 1.0, 0.0]
[root |INFO|deepbiome.py:199] Compute time : 3.6023569107055664
[root |INFO|deepbiome.py:200] 1 fold computing end!---------------------------------------------
[root |INFO|deepbiome.py:153] -------2 simulation start!----------------------------------
[root |INFO|readers.py:58] -----------------------------------------------------------------------
[root |INFO|readers.py:59] Construct Dataset
[root |INFO|readers.py:60] -----------------------------------------------------------------------
[root |INFO|readers.py:61] Load data
[root |INFO|deepbiome.py:164] -----------------------------------------------------------------
[root |INFO|deepbiome.py:165] Build network for 2 simulation
[root |INFO|build_network.py:521] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:522] Read phylogenetic tree information from /DATA/home/muha/github_repos/deepbiome/deepbiome/tests/data/genus48_dic.csv
[root |INFO|build_network.py:528] Phylogenetic tree level list: ['Genus', 'Family', 'Order', 'Class', 'Phylum']
[root |INFO|build_network.py:529] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:537] Genus: 48
[root |INFO|build_network.py:537] Family: 40
[root |INFO|build_network.py:537] Order: 23
[root |INFO|build_network.py:537] Class: 17
[root |INFO|build_network.py:537] Phylum: 9
[root |INFO|build_network.py:543] Phylum_with_covariates: 17
[root |INFO|build_network.py:546] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:547] Phylogenetic_tree_dict info: ['Genus', 'Order', 'Class', 'Family', 'Phylum_with_covariates', 'Phylum', 'Number']
[root |INFO|build_network.py:548] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:558] Build edge weights between [ Genus, Family]
[root |INFO|build_network.py:558] Build edge weights between [Family, Order]
[root |INFO|build_network.py:558] Build edge weights between [ Order, Class]
[root |INFO|build_network.py:558] Build edge weights between [ Class, Phylum]
[root |INFO|build_network.py:571] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:586] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:587] Build network based on phylogenetic tree information
[root |INFO|build_network.py:588] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:670] ------------------------------------------------------------------------------------------
Model: "model_1"
__________________________________________________________________________________________________
Layer (type) Output Shape Param # Connected to
==================================================================================================
input (InputLayer) (None, 48) 0
__________________________________________________________________________________________________
l1_dense (Dense_with_tree) (None, 40) 1960 input[0][0]
__________________________________________________________________________________________________
l1_activation (Activation) (None, 40) 0 l1_dense[0][0]
__________________________________________________________________________________________________
l2_dense (Dense_with_tree) (None, 23) 943 l1_activation[0][0]
__________________________________________________________________________________________________
l2_activation (Activation) (None, 23) 0 l2_dense[0][0]
__________________________________________________________________________________________________
l3_dense (Dense_with_tree) (None, 17) 408 l2_activation[0][0]
__________________________________________________________________________________________________
l3_activation (Activation) (None, 17) 0 l3_dense[0][0]
__________________________________________________________________________________________________
l4_dense (Dense_with_tree) (None, 9) 162 l3_activation[0][0]
__________________________________________________________________________________________________
l4_activation (Activation) (None, 9) 0 l4_dense[0][0]
__________________________________________________________________________________________________
covariates_input (InputLayer) (None, 8) 0
__________________________________________________________________________________________________
biome_covariates_concat (Concat (None, 17) 0 l4_activation[0][0]
covariates_input[0][0]
__________________________________________________________________________________________________
last_dense_h (Dense) (None, 1) 18 biome_covariates_concat[0][0]
__________________________________________________________________________________________________
p_hat (Activation) (None, 1) 0 last_dense_h[0][0]
==================================================================================================
Total params: 3,491
Trainable params: 3,491
Non-trainable params: 0
__________________________________________________________________________________________________
[root |INFO|build_network.py:61] Build Network
[root |INFO|build_network.py:62] Optimizer = adam
[root |INFO|build_network.py:63] Loss = binary_crossentropy
[root |INFO|build_network.py:64] Metrics = binary_accuracy, sensitivity, specificity, gmeasure
[root |INFO|deepbiome.py:176] -----------------------------------------------------------------
[root |INFO|deepbiome.py:177] 2 fold computing start!----------------------------------
[root |INFO|build_network.py:137] Training start!
Train on 640 samples, validate on 160 samples
Epoch 1/10
640/640 [==============================] - 1s 782us/step - loss: 2.5421 - binary_accuracy: 0.5859 - sensitivity: 0.7057 - specificity: 0.2490 - gmeasure: 0.3932 - val_loss: 1.6270 - val_binary_accuracy: 0.6250 - val_sensitivity: 0.7962 - val_specificity: 0.2233 - val_gmeasure: 0.4150
Epoch 2/10
640/640 [==============================] - 0s 116us/step - loss: 1.9945 - binary_accuracy: 0.5688 - sensitivity: 0.6823 - specificity: 0.2938 - gmeasure: 0.4414 - val_loss: 1.2290 - val_binary_accuracy: 0.6062 - val_sensitivity: 0.7127 - val_specificity: 0.3178 - val_gmeasure: 0.4747
Epoch 3/10
640/640 [==============================] - 0s 115us/step - loss: 1.4228 - binary_accuracy: 0.5906 - sensitivity: 0.7194 - specificity: 0.2600 - gmeasure: 0.4167 - val_loss: 0.8595 - val_binary_accuracy: 0.6438 - val_sensitivity: 0.8164 - val_specificity: 0.2233 - val_gmeasure: 0.4195
Epoch 4/10
640/640 [==============================] - 0s 110us/step - loss: 0.9308 - binary_accuracy: 0.5922 - sensitivity: 0.7519 - specificity: 0.1935 - gmeasure: 0.3590 - val_loss: 0.6228 - val_binary_accuracy: 0.6938 - val_sensitivity: 0.8991 - val_specificity: 0.2233 - val_gmeasure: 0.4411
Epoch 5/10
640/640 [==============================] - 0s 110us/step - loss: 0.6570 - binary_accuracy: 0.6703 - sensitivity: 0.9030 - specificity: 0.1052 - gmeasure: 0.1911 - val_loss: 0.6004 - val_binary_accuracy: 0.7188 - val_sensitivity: 0.9793 - val_specificity: 0.0000e+00 - val_gmeasure: 0.0000e+00
Epoch 6/10
640/640 [==============================] - 0s 112us/step - loss: 0.6270 - binary_accuracy: 0.7031 - sensitivity: 0.9651 - specificity: 0.0457 - gmeasure: 0.1232 - val_loss: 0.6574 - val_binary_accuracy: 0.6000 - val_sensitivity: 0.7574 - val_specificity: 0.1707 - val_gmeasure: 0.3037
Epoch 7/10
640/640 [==============================] - 0s 114us/step - loss: 0.6275 - binary_accuracy: 0.6844 - sensitivity: 0.9296 - specificity: 0.0619 - gmeasure: 0.1349 - val_loss: 0.6251 - val_binary_accuracy: 0.7188 - val_sensitivity: 0.9512 - val_specificity: 0.1480 - val_gmeasure: 0.3174
Epoch 8/10
640/640 [==============================] - 0s 120us/step - loss: 0.6198 - binary_accuracy: 0.7078 - sensitivity: 0.9500 - specificity: 0.0928 - gmeasure: 0.1620 - val_loss: 0.5917 - val_binary_accuracy: 0.7375 - val_sensitivity: 1.0000 - val_specificity: 0.0000e+00 - val_gmeasure: 0.0000e+00
Epoch 9/10
640/640 [==============================] - 0s 113us/step - loss: 0.6121 - binary_accuracy: 0.7094 - sensitivity: 0.9925 - specificity: 0.0000e+00 - gmeasure: 0.0000e+00 - val_loss: 0.5936 - val_binary_accuracy: 0.7375 - val_sensitivity: 1.0000 - val_specificity: 0.0000e+00 - val_gmeasure: 0.0000e+00
Epoch 10/10
640/640 [==============================] - 0s 118us/step - loss: 0.6053 - binary_accuracy: 0.7156 - sensitivity: 1.0000 - specificity: 0.0000e+00 - gmeasure: 0.0000e+00 - val_loss: 0.5944 - val_binary_accuracy: 0.7375 - val_sensitivity: 1.0000 - val_specificity: 0.0000e+00 - val_gmeasure: 0.0000e+00
[root |INFO|build_network.py:87] Load trained model weight at ./example_result/weight_1.h5
[root |INFO|build_network.py:147] Training end with time 2.6916720867156982!
[root |INFO|build_network.py:83] Saved trained model weight at ./example_result/weight_1.h5
[root |DEBUG|deepbiome.py:185] Save weight at ./example_result/weight_1.h5
[root |DEBUG|deepbiome.py:188] Save history at ./example_result/hist_1.json
[root |INFO|build_network.py:173] Evaluation start!
800/800 [==============================] - 0s 3us/step
[root |INFO|build_network.py:178] Evaluation end with time 0.008504152297973633!
[root |INFO|build_network.py:179] Evaluation: [0.6034644246101379, 0.7200000286102295, 1.0, 0.0, 0.0]
[root |INFO|build_network.py:173] Evaluation start!
200/200 [==============================] - 0s 9us/step
[root |INFO|build_network.py:178] Evaluation end with time 0.007249355316162109!
[root |INFO|build_network.py:179] Evaluation: [0.6046146154403687, 0.7200000286102295, 1.0, 0.0, 0.0]
[root |INFO|deepbiome.py:199] Compute time : 3.215890407562256
[root |INFO|deepbiome.py:200] 2 fold computing end!---------------------------------------------
[root |INFO|deepbiome.py:153] -------3 simulation start!----------------------------------
[root |INFO|readers.py:58] -----------------------------------------------------------------------
[root |INFO|readers.py:59] Construct Dataset
[root |INFO|readers.py:60] -----------------------------------------------------------------------
[root |INFO|readers.py:61] Load data
[root |INFO|deepbiome.py:164] -----------------------------------------------------------------
[root |INFO|deepbiome.py:165] Build network for 3 simulation
[root |INFO|build_network.py:521] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:522] Read phylogenetic tree information from /DATA/home/muha/github_repos/deepbiome/deepbiome/tests/data/genus48_dic.csv
[root |INFO|build_network.py:528] Phylogenetic tree level list: ['Genus', 'Family', 'Order', 'Class', 'Phylum']
[root |INFO|build_network.py:529] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:537] Genus: 48
[root |INFO|build_network.py:537] Family: 40
[root |INFO|build_network.py:537] Order: 23
[root |INFO|build_network.py:537] Class: 17
[root |INFO|build_network.py:537] Phylum: 9
[root |INFO|build_network.py:543] Phylum_with_covariates: 17
[root |INFO|build_network.py:546] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:547] Phylogenetic_tree_dict info: ['Genus', 'Order', 'Class', 'Family', 'Phylum_with_covariates', 'Phylum', 'Number']
[root |INFO|build_network.py:548] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:558] Build edge weights between [ Genus, Family]
[root |INFO|build_network.py:558] Build edge weights between [Family, Order]
[root |INFO|build_network.py:558] Build edge weights between [ Order, Class]
[root |INFO|build_network.py:558] Build edge weights between [ Class, Phylum]
[root |INFO|build_network.py:571] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:586] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:587] Build network based on phylogenetic tree information
[root |INFO|build_network.py:588] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:670] ------------------------------------------------------------------------------------------
Model: "model_1"
__________________________________________________________________________________________________
Layer (type) Output Shape Param # Connected to
==================================================================================================
input (InputLayer) (None, 48) 0
__________________________________________________________________________________________________
l1_dense (Dense_with_tree) (None, 40) 1960 input[0][0]
__________________________________________________________________________________________________
l1_activation (Activation) (None, 40) 0 l1_dense[0][0]
__________________________________________________________________________________________________
l2_dense (Dense_with_tree) (None, 23) 943 l1_activation[0][0]
__________________________________________________________________________________________________
l2_activation (Activation) (None, 23) 0 l2_dense[0][0]
__________________________________________________________________________________________________
l3_dense (Dense_with_tree) (None, 17) 408 l2_activation[0][0]
__________________________________________________________________________________________________
l3_activation (Activation) (None, 17) 0 l3_dense[0][0]
__________________________________________________________________________________________________
l4_dense (Dense_with_tree) (None, 9) 162 l3_activation[0][0]
__________________________________________________________________________________________________
l4_activation (Activation) (None, 9) 0 l4_dense[0][0]
__________________________________________________________________________________________________
covariates_input (InputLayer) (None, 8) 0
__________________________________________________________________________________________________
biome_covariates_concat (Concat (None, 17) 0 l4_activation[0][0]
covariates_input[0][0]
__________________________________________________________________________________________________
last_dense_h (Dense) (None, 1) 18 biome_covariates_concat[0][0]
__________________________________________________________________________________________________
p_hat (Activation) (None, 1) 0 last_dense_h[0][0]
==================================================================================================
Total params: 3,491
Trainable params: 3,491
Non-trainable params: 0
__________________________________________________________________________________________________
[root |INFO|build_network.py:61] Build Network
[root |INFO|build_network.py:62] Optimizer = adam
[root |INFO|build_network.py:63] Loss = binary_crossentropy
[root |INFO|build_network.py:64] Metrics = binary_accuracy, sensitivity, specificity, gmeasure
[root |INFO|deepbiome.py:176] -----------------------------------------------------------------
[root |INFO|deepbiome.py:177] 3 fold computing start!----------------------------------
[root |INFO|build_network.py:137] Training start!
Train on 640 samples, validate on 160 samples
Epoch 1/10
640/640 [==============================] - 0s 698us/step - loss: 4.9752 - binary_accuracy: 0.3828 - sensitivity: 0.2109 - specificity: 0.8032 - gmeasure: 0.2862 - val_loss: 1.6229 - val_binary_accuracy: 0.5813 - val_sensitivity: 0.6279 - val_specificity: 0.3807 - val_gmeasure: 0.4665
Epoch 2/10
640/640 [==============================] - 0s 105us/step - loss: 1.8461 - binary_accuracy: 0.6562 - sensitivity: 0.8684 - specificity: 0.1641 - gmeasure: 0.3599 - val_loss: 1.8489 - val_binary_accuracy: 0.6812 - val_sensitivity: 0.9384 - val_specificity: 0.1234 - val_gmeasure: 0.2155
Epoch 3/10
640/640 [==============================] - 0s 96us/step - loss: 1.6691 - binary_accuracy: 0.6375 - sensitivity: 0.8160 - specificity: 0.2189 - gmeasure: 0.3934 - val_loss: 1.4692 - val_binary_accuracy: 0.5625 - val_sensitivity: 0.6009 - val_specificity: 0.4016 - val_gmeasure: 0.4711
Epoch 4/10
640/640 [==============================] - 0s 116us/step - loss: 1.4963 - binary_accuracy: 0.5781 - sensitivity: 0.6485 - specificity: 0.4319 - gmeasure: 0.5228 - val_loss: 1.2639 - val_binary_accuracy: 0.6375 - val_sensitivity: 0.7273 - val_specificity: 0.2557 - val_gmeasure: 0.3777
Epoch 5/10
640/640 [==============================] - 0s 104us/step - loss: 1.2753 - binary_accuracy: 0.6313 - sensitivity: 0.8003 - specificity: 0.2395 - gmeasure: 0.4309 - val_loss: 1.0925 - val_binary_accuracy: 0.6438 - val_sensitivity: 0.7585 - val_specificity: 0.2557 - val_gmeasure: 0.3777
Epoch 6/10
640/640 [==============================] - 0s 88us/step - loss: 1.0665 - binary_accuracy: 0.6187 - sensitivity: 0.7677 - specificity: 0.2748 - gmeasure: 0.4283 - val_loss: 0.9395 - val_binary_accuracy: 0.6187 - val_sensitivity: 0.6985 - val_specificity: 0.3807 - val_gmeasure: 0.4942
Epoch 7/10
640/640 [==============================] - 0s 94us/step - loss: 0.9074 - binary_accuracy: 0.6094 - sensitivity: 0.7371 - specificity: 0.3136 - gmeasure: 0.4668 - val_loss: 0.7888 - val_binary_accuracy: 0.6812 - val_sensitivity: 0.9099 - val_specificity: 0.1799 - val_gmeasure: 0.3147
Epoch 8/10
640/640 [==============================] - 0s 113us/step - loss: 0.7632 - binary_accuracy: 0.6391 - sensitivity: 0.8310 - specificity: 0.1965 - gmeasure: 0.3900 - val_loss: 0.7070 - val_binary_accuracy: 0.6187 - val_sensitivity: 0.6813 - val_specificity: 0.4135 - val_gmeasure: 0.5097
Epoch 9/10
640/640 [==============================] - 0s 112us/step - loss: 0.6805 - binary_accuracy: 0.6531 - sensitivity: 0.8495 - specificity: 0.2153 - gmeasure: 0.4098 - val_loss: 0.6269 - val_binary_accuracy: 0.6750 - val_sensitivity: 0.8896 - val_specificity: 0.2022 - val_gmeasure: 0.3508
Epoch 10/10
640/640 [==============================] - 0s 121us/step - loss: 0.6260 - binary_accuracy: 0.6938 - sensitivity: 0.9586 - specificity: 0.0907 - gmeasure: 0.2173 - val_loss: 0.6025 - val_binary_accuracy: 0.7063 - val_sensitivity: 0.9782 - val_specificity: 0.0655 - val_gmeasure: 0.1751
[root |INFO|build_network.py:87] Load trained model weight at ./example_result/weight_2.h5
[root |INFO|build_network.py:147] Training end with time 2.9351305961608887!
[root |INFO|build_network.py:83] Saved trained model weight at ./example_result/weight_2.h5
[root |DEBUG|deepbiome.py:185] Save weight at ./example_result/weight_2.h5
[root |DEBUG|deepbiome.py:188] Save history at ./example_result/hist_2.json
[root |INFO|build_network.py:173] Evaluation start!
800/800 [==============================] - 0s 3us/step
[root |INFO|build_network.py:178] Evaluation end with time 0.0087738037109375!
[root |INFO|build_network.py:179] Evaluation: [0.6114577054977417, 0.6949999928474426, 0.9731183052062988, 0.05371900647878647, 0.22863715887069702]
[root |INFO|build_network.py:173] Evaluation start!
200/200 [==============================] - 0s 41us/step
[root |INFO|build_network.py:178] Evaluation end with time 0.014694690704345703!
[root |INFO|build_network.py:179] Evaluation: [0.6872313022613525, 0.6100000143051147, 0.9312977194786072, 0.0, 0.0]
[root |INFO|deepbiome.py:199] Compute time : 3.4203402996063232
[root |INFO|deepbiome.py:200] 3 fold computing end!---------------------------------------------
[root |INFO|deepbiome.py:153] -------4 simulation start!----------------------------------
[root |INFO|readers.py:58] -----------------------------------------------------------------------
[root |INFO|readers.py:59] Construct Dataset
[root |INFO|readers.py:60] -----------------------------------------------------------------------
[root |INFO|readers.py:61] Load data
[root |INFO|deepbiome.py:164] -----------------------------------------------------------------
[root |INFO|deepbiome.py:165] Build network for 4 simulation
[root |INFO|build_network.py:521] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:522] Read phylogenetic tree information from /DATA/home/muha/github_repos/deepbiome/deepbiome/tests/data/genus48_dic.csv
[root |INFO|build_network.py:528] Phylogenetic tree level list: ['Genus', 'Family', 'Order', 'Class', 'Phylum']
[root |INFO|build_network.py:529] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:537] Genus: 48
[root |INFO|build_network.py:537] Family: 40
[root |INFO|build_network.py:537] Order: 23
[root |INFO|build_network.py:537] Class: 17
[root |INFO|build_network.py:537] Phylum: 9
[root |INFO|build_network.py:543] Phylum_with_covariates: 17
[root |INFO|build_network.py:546] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:547] Phylogenetic_tree_dict info: ['Genus', 'Order', 'Class', 'Family', 'Phylum_with_covariates', 'Phylum', 'Number']
[root |INFO|build_network.py:548] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:558] Build edge weights between [ Genus, Family]
[root |INFO|build_network.py:558] Build edge weights between [Family, Order]
[root |INFO|build_network.py:558] Build edge weights between [ Order, Class]
[root |INFO|build_network.py:558] Build edge weights between [ Class, Phylum]
[root |INFO|build_network.py:571] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:586] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:587] Build network based on phylogenetic tree information
[root |INFO|build_network.py:588] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:670] ------------------------------------------------------------------------------------------
Model: "model_1"
__________________________________________________________________________________________________
Layer (type) Output Shape Param # Connected to
==================================================================================================
input (InputLayer) (None, 48) 0
__________________________________________________________________________________________________
l1_dense (Dense_with_tree) (None, 40) 1960 input[0][0]
__________________________________________________________________________________________________
l1_activation (Activation) (None, 40) 0 l1_dense[0][0]
__________________________________________________________________________________________________
l2_dense (Dense_with_tree) (None, 23) 943 l1_activation[0][0]
__________________________________________________________________________________________________
l2_activation (Activation) (None, 23) 0 l2_dense[0][0]
__________________________________________________________________________________________________
l3_dense (Dense_with_tree) (None, 17) 408 l2_activation[0][0]
__________________________________________________________________________________________________
l3_activation (Activation) (None, 17) 0 l3_dense[0][0]
__________________________________________________________________________________________________
l4_dense (Dense_with_tree) (None, 9) 162 l3_activation[0][0]
__________________________________________________________________________________________________
l4_activation (Activation) (None, 9) 0 l4_dense[0][0]
__________________________________________________________________________________________________
covariates_input (InputLayer) (None, 8) 0
__________________________________________________________________________________________________
biome_covariates_concat (Concat (None, 17) 0 l4_activation[0][0]
covariates_input[0][0]
__________________________________________________________________________________________________
last_dense_h (Dense) (None, 1) 18 biome_covariates_concat[0][0]
__________________________________________________________________________________________________
p_hat (Activation) (None, 1) 0 last_dense_h[0][0]
==================================================================================================
Total params: 3,491
Trainable params: 3,491
Non-trainable params: 0
__________________________________________________________________________________________________
[root |INFO|build_network.py:61] Build Network
[root |INFO|build_network.py:62] Optimizer = adam
[root |INFO|build_network.py:63] Loss = binary_crossentropy
[root |INFO|build_network.py:64] Metrics = binary_accuracy, sensitivity, specificity, gmeasure
[root |INFO|deepbiome.py:176] -----------------------------------------------------------------
[root |INFO|deepbiome.py:177] 4 fold computing start!----------------------------------
[root |INFO|build_network.py:137] Training start!
Train on 640 samples, validate on 160 samples
Epoch 1/10
640/640 [==============================] - 1s 785us/step - loss: 0.8356 - binary_accuracy: 0.5906 - sensitivity: 0.7687 - specificity: 0.2441 - gmeasure: 0.3609 - val_loss: 0.6479 - val_binary_accuracy: 0.6875 - val_sensitivity: 0.9545 - val_specificity: 0.0882 - val_gmeasure: 0.2000
Epoch 2/10
640/640 [==============================] - 0s 377us/step - loss: 0.6636 - binary_accuracy: 0.6625 - sensitivity: 0.9362 - specificity: 0.0742 - gmeasure: 0.1698 - val_loss: 0.6219 - val_binary_accuracy: 0.6875 - val_sensitivity: 1.0000 - val_specificity: 0.0000e+00 - val_gmeasure: 0.0000e+00
Epoch 3/10
640/640 [==============================] - 0s 436us/step - loss: 0.6364 - binary_accuracy: 0.6875 - sensitivity: 1.0000 - specificity: 0.0059 - gmeasure: 0.0213 - val_loss: 0.6247 - val_binary_accuracy: 0.6875 - val_sensitivity: 1.0000 - val_specificity: 0.0000e+00 - val_gmeasure: 0.0000e+00
Epoch 4/10
640/640 [==============================] - 0s 115us/step - loss: 0.6366 - binary_accuracy: 0.6891 - sensitivity: 1.0000 - specificity: 0.0081 - gmeasure: 0.0250 - val_loss: 0.6417 - val_binary_accuracy: 0.6875 - val_sensitivity: 1.0000 - val_specificity: 0.0000e+00 - val_gmeasure: 0.0000e+00
Epoch 5/10
640/640 [==============================] - 0s 204us/step - loss: 0.6337 - binary_accuracy: 0.6875 - sensitivity: 1.0000 - specificity: 0.0070 - gmeasure: 0.0232 - val_loss: 0.6225 - val_binary_accuracy: 0.6875 - val_sensitivity: 1.0000 - val_specificity: 0.0000e+00 - val_gmeasure: 0.0000e+00
Epoch 6/10
640/640 [==============================] - 0s 440us/step - loss: 0.6337 - binary_accuracy: 0.6797 - sensitivity: 0.9909 - specificity: 0.0000e+00 - gmeasure: 0.0000e+00 - val_loss: 0.6272 - val_binary_accuracy: 0.6875 - val_sensitivity: 1.0000 - val_specificity: 0.0000e+00 - val_gmeasure: 0.0000e+00
Epoch 7/10
640/640 [==============================] - 0s 298us/step - loss: 0.6380 - binary_accuracy: 0.6859 - sensitivity: 1.0000 - specificity: 0.0000e+00 - gmeasure: 0.0000e+00 - val_loss: 0.6209 - val_binary_accuracy: 0.6875 - val_sensitivity: 1.0000 - val_specificity: 0.0000e+00 - val_gmeasure: 0.0000e+00
Epoch 8/10
640/640 [==============================] - 0s 103us/step - loss: 0.6307 - binary_accuracy: 0.6844 - sensitivity: 0.9978 - specificity: 0.0000e+00 - gmeasure: 0.0000e+00 - val_loss: 0.6168 - val_binary_accuracy: 0.6875 - val_sensitivity: 1.0000 - val_specificity: 0.0000e+00 - val_gmeasure: 0.0000e+00
Epoch 9/10
640/640 [==============================] - 0s 95us/step - loss: 0.6273 - binary_accuracy: 0.6875 - sensitivity: 1.0000 - specificity: 0.0059 - gmeasure: 0.0213 - val_loss: 0.6196 - val_binary_accuracy: 0.6938 - val_sensitivity: 1.0000 - val_specificity: 0.0147 - val_gmeasure: 0.0606
Epoch 10/10
640/640 [==============================] - 0s 96us/step - loss: 0.6246 - binary_accuracy: 0.6875 - sensitivity: 1.0000 - specificity: 0.0051 - gmeasure: 0.0199 - val_loss: 0.6234 - val_binary_accuracy: 0.6875 - val_sensitivity: 0.9934 - val_specificity: 0.0147 - val_gmeasure: 0.0606
[root |INFO|build_network.py:87] Load trained model weight at ./example_result/weight_3.h5
[root |INFO|build_network.py:147] Training end with time 3.7450311183929443!
[root |INFO|build_network.py:83] Saved trained model weight at ./example_result/weight_3.h5
[root |DEBUG|deepbiome.py:185] Save weight at ./example_result/weight_3.h5
[root |DEBUG|deepbiome.py:188] Save history at ./example_result/hist_3.json
[root |INFO|build_network.py:173] Evaluation start!
800/800 [==============================] - 0s 4us/step
[root |INFO|build_network.py:178] Evaluation end with time 0.011616230010986328!
[root |INFO|build_network.py:179] Evaluation: [0.6179130673408508, 0.6862499713897705, 1.0, 0.0, 0.0]
[root |INFO|build_network.py:173] Evaluation start!
200/200 [==============================] - 0s 11us/step
[root |INFO|build_network.py:178] Evaluation end with time 0.0076541900634765625!
[root |INFO|build_network.py:179] Evaluation: [0.6171166300773621, 0.6899999976158142, 1.0, 0.0, 0.0]
[root |INFO|deepbiome.py:199] Compute time : 4.296782732009888
[root |INFO|deepbiome.py:200] 4 fold computing end!---------------------------------------------
[root |INFO|deepbiome.py:153] -------5 simulation start!----------------------------------
[root |INFO|readers.py:58] -----------------------------------------------------------------------
[root |INFO|readers.py:59] Construct Dataset
[root |INFO|readers.py:60] -----------------------------------------------------------------------
[root |INFO|readers.py:61] Load data
[root |INFO|deepbiome.py:164] -----------------------------------------------------------------
[root |INFO|deepbiome.py:165] Build network for 5 simulation
[root |INFO|build_network.py:521] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:522] Read phylogenetic tree information from /DATA/home/muha/github_repos/deepbiome/deepbiome/tests/data/genus48_dic.csv
[root |INFO|build_network.py:528] Phylogenetic tree level list: ['Genus', 'Family', 'Order', 'Class', 'Phylum']
[root |INFO|build_network.py:529] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:537] Genus: 48
[root |INFO|build_network.py:537] Family: 40
[root |INFO|build_network.py:537] Order: 23
[root |INFO|build_network.py:537] Class: 17
[root |INFO|build_network.py:537] Phylum: 9
[root |INFO|build_network.py:543] Phylum_with_covariates: 17
[root |INFO|build_network.py:546] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:547] Phylogenetic_tree_dict info: ['Genus', 'Order', 'Class', 'Family', 'Phylum_with_covariates', 'Phylum', 'Number']
[root |INFO|build_network.py:548] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:558] Build edge weights between [ Genus, Family]
[root |INFO|build_network.py:558] Build edge weights between [Family, Order]
[root |INFO|build_network.py:558] Build edge weights between [ Order, Class]
[root |INFO|build_network.py:558] Build edge weights between [ Class, Phylum]
[root |INFO|build_network.py:571] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:586] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:587] Build network based on phylogenetic tree information
[root |INFO|build_network.py:588] ------------------------------------------------------------------------------------------
[root |INFO|build_network.py:670] ------------------------------------------------------------------------------------------
Model: "model_1"
__________________________________________________________________________________________________
Layer (type) Output Shape Param # Connected to
==================================================================================================
input (InputLayer) (None, 48) 0
__________________________________________________________________________________________________
l1_dense (Dense_with_tree) (None, 40) 1960 input[0][0]
__________________________________________________________________________________________________
l1_activation (Activation) (None, 40) 0 l1_dense[0][0]
__________________________________________________________________________________________________
l2_dense (Dense_with_tree) (None, 23) 943 l1_activation[0][0]
__________________________________________________________________________________________________
l2_activation (Activation) (None, 23) 0 l2_dense[0][0]
__________________________________________________________________________________________________
l3_dense (Dense_with_tree) (None, 17) 408 l2_activation[0][0]
__________________________________________________________________________________________________
l3_activation (Activation) (None, 17) 0 l3_dense[0][0]
__________________________________________________________________________________________________
l4_dense (Dense_with_tree) (None, 9) 162 l3_activation[0][0]
__________________________________________________________________________________________________
l4_activation (Activation) (None, 9) 0 l4_dense[0][0]
__________________________________________________________________________________________________
covariates_input (InputLayer) (None, 8) 0
__________________________________________________________________________________________________
biome_covariates_concat (Concat (None, 17) 0 l4_activation[0][0]
covariates_input[0][0]
__________________________________________________________________________________________________
last_dense_h (Dense) (None, 1) 18 biome_covariates_concat[0][0]
__________________________________________________________________________________________________
p_hat (Activation) (None, 1) 0 last_dense_h[0][0]
==================================================================================================
Total params: 3,491
Trainable params: 3,491
Non-trainable params: 0
__________________________________________________________________________________________________
[root |INFO|build_network.py:61] Build Network
[root |INFO|build_network.py:62] Optimizer = adam
[root |INFO|build_network.py:63] Loss = binary_crossentropy
[root |INFO|build_network.py:64] Metrics = binary_accuracy, sensitivity, specificity, gmeasure
[root |INFO|deepbiome.py:176] -----------------------------------------------------------------
[root |INFO|deepbiome.py:177] 5 fold computing start!----------------------------------
[root |INFO|build_network.py:137] Training start!
Train on 640 samples, validate on 160 samples
Epoch 1/10
640/640 [==============================] - 0s 712us/step - loss: 2.8599 - binary_accuracy: 0.6828 - sensitivity: 0.9640 - specificity: 0.0585 - gmeasure: 0.1176 - val_loss: 1.2346 - val_binary_accuracy: 0.5437 - val_sensitivity: 0.6156 - val_specificity: 0.3758 - val_gmeasure: 0.4799
Epoch 2/10
640/640 [==============================] - 0s 127us/step - loss: 1.5378 - binary_accuracy: 0.4844 - sensitivity: 0.4592 - specificity: 0.5396 - gmeasure: 0.4676 - val_loss: 1.1423 - val_binary_accuracy: 0.6187 - val_sensitivity: 0.8014 - val_specificity: 0.2035 - val_gmeasure: 0.4001
Epoch 3/10
640/640 [==============================] - 0s 152us/step - loss: 1.2258 - binary_accuracy: 0.6359 - sensitivity: 0.8343 - specificity: 0.2156 - gmeasure: 0.4089 - val_loss: 0.9623 - val_binary_accuracy: 0.5938 - val_sensitivity: 0.7371 - val_specificity: 0.3243 - val_gmeasure: 0.4863
Epoch 4/10
640/640 [==============================] - 0s 140us/step - loss: 0.9657 - binary_accuracy: 0.5641 - sensitivity: 0.6153 - specificity: 0.4465 - gmeasure: 0.5166 - val_loss: 0.8278 - val_binary_accuracy: 0.6313 - val_sensitivity: 0.8227 - val_specificity: 0.1879 - val_gmeasure: 0.3905
Epoch 5/10
640/640 [==============================] - 0s 129us/step - loss: 0.8022 - binary_accuracy: 0.6297 - sensitivity: 0.8065 - specificity: 0.2489 - gmeasure: 0.4133 - val_loss: 0.7336 - val_binary_accuracy: 0.5500 - val_sensitivity: 0.6303 - val_specificity: 0.3555 - val_gmeasure: 0.4700
Epoch 6/10
640/640 [==============================] - 0s 142us/step - loss: 0.6674 - binary_accuracy: 0.6375 - sensitivity: 0.7970 - specificity: 0.2447 - gmeasure: 0.3655 - val_loss: 0.6640 - val_binary_accuracy: 0.7063 - val_sensitivity: 1.0000 - val_specificity: 0.0312 - val_gmeasure: 0.0884
Epoch 7/10
640/640 [==============================] - 0s 148us/step - loss: 0.6512 - binary_accuracy: 0.6578 - sensitivity: 0.8994 - specificity: 0.1401 - gmeasure: 0.2796 - val_loss: 0.6310 - val_binary_accuracy: 0.6625 - val_sensitivity: 0.9004 - val_specificity: 0.0469 - val_gmeasure: 0.1050
Epoch 8/10
640/640 [==============================] - 0s 130us/step - loss: 0.6343 - binary_accuracy: 0.6797 - sensitivity: 0.9750 - specificity: 0.0571 - gmeasure: 0.1793 - val_loss: 0.6247 - val_binary_accuracy: 0.6812 - val_sensitivity: 0.9216 - val_specificity: 0.0469 - val_gmeasure: 0.1066
Epoch 9/10
640/640 [==============================] - 0s 130us/step - loss: 0.6340 - binary_accuracy: 0.6719 - sensitivity: 0.9664 - specificity: 0.0424 - gmeasure: 0.1189 - val_loss: 0.6255 - val_binary_accuracy: 0.7000 - val_sensitivity: 0.9929 - val_specificity: 0.0312 - val_gmeasure: 0.0884
Epoch 10/10
640/640 [==============================] - 0s 132us/step - loss: 0.6359 - binary_accuracy: 0.6703 - sensitivity: 0.9687 - specificity: 0.0390 - gmeasure: 0.1287 - val_loss: 0.6267 - val_binary_accuracy: 0.6687 - val_sensitivity: 0.9071 - val_specificity: 0.0469 - val_gmeasure: 0.1050
[root |INFO|build_network.py:87] Load trained model weight at ./example_result/weight_4.h5
[root |INFO|build_network.py:147] Training end with time 2.841571569442749!
[root |INFO|build_network.py:83] Saved trained model weight at ./example_result/weight_4.h5
[root |DEBUG|deepbiome.py:185] Save weight at ./example_result/weight_4.h5
[root |DEBUG|deepbiome.py:188] Save history at ./example_result/hist_4.json
[root |INFO|build_network.py:173] Evaluation start!
800/800 [==============================] - 0s 9us/step
[root |INFO|build_network.py:178] Evaluation end with time 0.0183563232421875!
[root |INFO|build_network.py:179] Evaluation: [0.6290234327316284, 0.6800000071525574, 0.9798534512519836, 0.035433072596788406, 0.18633092939853668]
[root |INFO|build_network.py:173] Evaluation start!
200/200 [==============================] - 0s 37us/step
[root |INFO|build_network.py:178] Evaluation end with time 0.016957521438598633!
[root |INFO|build_network.py:179] Evaluation: [0.6027040481567383, 0.6949999928474426, 0.9716312289237976, 0.033898305147886276, 0.18148458003997803]
[root |INFO|deepbiome.py:199] Compute time : 3.4107158184051514
[root |INFO|deepbiome.py:200] 5 fold computing end!---------------------------------------------
[root |INFO|deepbiome.py:211] -----------------------------------------------------------------
[root |INFO|deepbiome.py:212] Train Evaluation : ['loss' 'binary_accuracy' 'sensitivity' 'specificity' 'gmeasure']
[root |INFO|deepbiome.py:213] mean : [2.705 0.619 0.791 0.218 0.083]
[root |INFO|deepbiome.py:214] std : [4.178 0.153 0.395 0.392 0.103]
[root |INFO|deepbiome.py:215] -----------------------------------------------------------------
[root |INFO|deepbiome.py:216] Test Evaluation : ['loss' 'binary_accuracy' 'sensitivity' 'specificity' 'gmeasure']
[root |INFO|deepbiome.py:217] mean : [2.727 0.605 0.781 0.207 0.036]
[root |INFO|deepbiome.py:218] std : [4.198 0.152 0.391 0.397 0.073]
[root |INFO|deepbiome.py:219] -----------------------------------------------------------------
[root |INFO|deepbiome.py:230] Total Computing Ended
[root |INFO|deepbiome.py:231] -----------------------------------------------------------------
The DeepBiome used the covariates at the latest hidden layer just before the last prediction layer (biome_covariates_concat layer in the above example).
[60]:
network_with_cov.model.summary()
Model: "model_1"
__________________________________________________________________________________________________
Layer (type) Output Shape Param # Connected to
==================================================================================================
input (InputLayer) (None, 48) 0
__________________________________________________________________________________________________
l1_dense (Dense_with_tree) (None, 40) 1960 input[0][0]
__________________________________________________________________________________________________
l1_activation (Activation) (None, 40) 0 l1_dense[0][0]
__________________________________________________________________________________________________
l2_dense (Dense_with_tree) (None, 23) 943 l1_activation[0][0]
__________________________________________________________________________________________________
l2_activation (Activation) (None, 23) 0 l2_dense[0][0]
__________________________________________________________________________________________________
l3_dense (Dense_with_tree) (None, 17) 408 l2_activation[0][0]
__________________________________________________________________________________________________
l3_activation (Activation) (None, 17) 0 l3_dense[0][0]
__________________________________________________________________________________________________
l4_dense (Dense_with_tree) (None, 9) 162 l3_activation[0][0]
__________________________________________________________________________________________________
l4_activation (Activation) (None, 9) 0 l4_dense[0][0]
__________________________________________________________________________________________________
covariates_input (InputLayer) (None, 8) 0
__________________________________________________________________________________________________
biome_covariates_concat (Concat (None, 17) 0 l4_activation[0][0]
covariates_input[0][0]
__________________________________________________________________________________________________
last_dense_h (Dense) (None, 1) 18 biome_covariates_concat[0][0]
__________________________________________________________________________________________________
p_hat (Activation) (None, 1) 0 last_dense_h[0][0]
==================================================================================================
Total params: 3,491
Trainable params: 3,491
Non-trainable params: 0
__________________________________________________________________________________________________
We can analysis the effects of covariates by checking the trained weights of the last hidden layer.
[61]:
weight_path = '%s/%s' % (prediction_path_info['model_info']['model_dir'], 'weight_0.h5')
trained_weight_list = deepbiome.deepbiome_get_trained_weight(log, network_info_with_covariates, path_info_with_covariates,
num_classes=1, weight_path=weight_path, verbose=False)
log.info(len(trained_weight_list))
[root |INFO|<ipython-input-61-b8f281eeb9fd>:4] 5
[62]:
trained_weight_list[4]
[62]:
| 0 | |
|---|---|
| Firmicutes | 0.566 |
| Actinobacteria | 0.330 |
| Bacteroidetes | -0.026 |
| Fusobacteria | 0.107 |
| Proteobacteria | 0.309 |
| TM7 | 0.340 |
| Spirochaetes | -0.235 |
| OD1 | 0.142 |
| Cyanobacteria_Chloroplast | -0.331 |
| Ages | -0.398 |
| Weights | -0.140 |
| Gender_1 | 0.019 |
| Ethnicity_1 | -0.114 |
| Ethnicity_2 | -0.202 |
| Ethnicity_3 | 0.487 |
| Ethnicity_4 | 0.411 |
| Ethnicity_5 | 0.691 |
Below is the phylogenetic tree plot for the network using covariates.
[63]:
phylum_color = ['lightsteelblue',
'moccasin',
'darkseagreen',
'khaki',
'mediumturquoise',
'lightblue',
'thistle',
'tan',
'mistyrose',
'honeydew',
'peachpuff',
'peru',
'saddlebrown',
'orchid',
'lightgray',
'gray',
'azure',
'lightcyan'
]
img = deepbiome.deepbiome_draw_phylogenetic_tree(log, network_info_with_covariates, path_info_with_covariates, num_classes=1,
file_name='%%inline', img_w=500, branch_vertical_margin=20,
arc_start=0, arc_span=360,
node_name_on=True, name_fsize=200,
tree_weight_on=True, tree_weight=trained_weight_list,
tree_level_list = ['Genus', 'Family', 'Order', 'Class', 'Phylum', 'Disease'],
weight_opacity=0.8, weight_max_radios=100,
phylum_background_color_on=True, phylum_color=phylum_color, phylum_color_legend=True,
show_covariates=True,
verbose=False)
img
[63]: